Abstract
Ecosystems are controlled by ‘bottom-up’ (resources) and ‘top-down’ (predation) forces. Viral infection is now recognized as a ubiquitous top-down control of microbial growth across ecosystems but, at the same time, cell death by viral predation influences, and is influenced by, resource availability. In this Review, we discuss recent advances in understanding the biogeochemical impact of viruses, focusing on how metabolic reprogramming of host cells during lytic viral infection alters the flow of energy and nutrients in aquatic ecosystems. Our synthesis revealed several emerging themes. First, viral infection transforms host metabolism, in part through virus-encoded metabolic genes; the functions performed by these genes appear to alleviate energetic and biosynthetic bottlenecks to viral production. Second, viral infection depends on the physiological state of the host cell and on environmental conditions, which are challenging to replicate in the laboratory. Last, metabolic reprogramming of infected cells and viral lysis alter nutrient cycling and carbon export in the oceans, although the net impacts remain uncertain. This Review highlights the need for understanding viral infection dynamics in realistic physiological and environmental contexts to better predict their biogeochemical consequences.
Similar content being viewed by others
Introduction
“A bacterium continually strives to produce two bacteria”, wrote François Jacob in The Logic of Life, “This seems to be its one project, its sole ambition” (ref.1). It is in relentless service of this goal that microorganisms collect substrates from their environment and assimilate them into new forms, harnessing energy from the Sun or chemical disequilibria, while competing with neighbours and relatives seeking to do the same. This globe-spanning effort has generated much of the biogeochemical structure in Earth’s environments: our oxygenated atmosphere; the widely nutrient-depleted surface ocean; and sulfide-rich and methane-rich sediments on land and sea2.
If microbial growth proceeded unchecked, however, global ecology would long ago have exhausted its resources and ground to a halt. Instead, death processes largely keep pace with growth in microbial ecosystems, and, in turn, drive nutrient recycling that fuels the growth of the next generation. In aquatic systems, a complex microbial food web of phytoplankton, heterotrophic bacteria and microscopic consumers was first proposed decades ago3. We now recognize additional mechanisms that are crucial for recycling biomass and supporting microbial growth, including lysis from viral infection4,5,6. During infection, the ‘sole ambition’ of the cell — to reproduce — is commandeered by the virus, which rewires host metabolism for the new goal of producing viral progeny7,8. As a result of this reprogramming, viral infection can have substantial biogeochemical consequences well before cellular lysis.
We can now appreciate that in virtually every microbial habitat on Earth, at any given time, there is a substantial portion of microbial cells whose ‘one project’ has been hijacked by viral infection4,9. Early efforts in metagenomics revealed the extensive novel genetic diversity of phages10,11, and subsequent work has vastly expanded our catalogue of diversity for both phages and archaeal viruses12,13,14,15. Assessment of eukaryotic viral diversity lags behind, in part due to the challenges of assembling whole eukaryotic viral genomes from marine field samples. Nevertheless, recent studies of eukaryotic viral diversity include genome sequences from cultured algal viruses that infect representatives of several different eukaryotic supergroups16,17,18,19, analysis of individual virally encoded genes20,21, metagenomic surveys from nature22 and targeted metagenomic assembly of uncultured viruses of protists23. Despite this rapid progress in genomics, our understanding of infection biology remains limited and is largely based on a few well-studied laboratory model systems; even less is known about infection dynamics in natural ecosystems. In this Review, we focus on cellular-level virus–host interactions and explore their potential impacts on biogeochemistry in aquatic systems. We first summarize how viruses alter the metabolism and composition of their host cells, with the caveat that much of this knowledge is based on a handful of model systems and laboratory growth conditions. We then summarize recent work exploring how host physiology and environmental conditions constrain or alter viral reprogramming of cellular pathways, cell lysis and viral production. Last, we extrapolate from these cellular-level processes to explore how viruses might impact biogeochemistry at the ecosystem scale.
Infection reprogrammes the cell
Viruses employ a range of infection strategies from lysogenic integration into the host genome to acute lytic bursting (Box 1); the prevalence of these strategies in the wild, and their physiological and environmental controls, are poorly understood. In this Review, we focus on lytic infection, the mode whose cellular and biogeochemical effects are best characterized. Lytic viral infection fundamentally reprogrammes host cell metabolism away from cellular replication towards progeny virus production (Fig. 1). This reprogramming is evident at the transcriptional level, where mRNA synthesis shifts rapidly and almost entirely to viral genes in a highly regulated manner. This transcriptional programme has been described for phages infecting cyanobacteria (cyanophages)24,25,26, phages of Cellulophaga baltica (marine Bacteroidetes)27,28 and viruses infecting diverse marine eukaryotic algae, including haptophytes29,30, prasinophytes31 and stramenopiles32. Comparative studies have found this gene expression programme to be relatively invariant for a given virus infecting different host strains of marine Synechococcus (Cyanobacteria)25 and C. baltica27,28, and for a given virus infecting host cells under phosphorus-replete and phosphorus-limited conditions31,33, although the onset of this transcriptional reprogramming can vary among individual cells in a population and may be stalled in non-growing or stationary phase cells34. Building on this knowledge from model systems, recent work has begun to detect spatiotemporal patterns of active infections in coastal and open ocean environments using metatranscriptomics35,36,37,38. Despite the relatively invariant viral gene expression programmes observed to date in cultures, the productivity of the infection can vary widely depending on host and environmental factors, as discussed below. Hence, a challenge going forward is to connect gene expression with other cellular markers of infection and viral production.
Schematic of selected metabolic fluxes towards biopolymer (nucleic acid and protein) production in a prototypical cyanobacterium, contrasting the metabolic states of an uninfected, replicating cell (part a) with a phage-infected, reprogrammed host cell (part b). Notable metabolic processes include proton pumping and electron transport by the photosystems in the thylakoid membranes; ATP synthesis; the interlocking carbon-fixing Calvin cycle and carbon-respiring pentose phosphate pathway; biosynthesis of nucleotides and amino acids; synthesis of DNA/RNA and proteins by polymerization; and pigment biosynthesis. These processes can lead to cell growth and replication, producing a daughter cell (part a), or lead to assembly of progeny phage particles and cell lysis (part b). In both cases, metabolic products are released to the environment, contributing to the dissolved organic matter (DOM) pool; this release is through exudation during growth (part a), whereas cell lysis releases cytoplasmic and membrane components of the killed host cell (part b). Orange arrows and the red cross (part b) indicate pathways stimulated or inhibited, respectively, during infection by expression of virally encoded accessory metabolic genes (note that many additional cellular pathways are affected or redirected by infection). TCA, tricarboxylic acid.
Beyond gene expression, lytic viral infection alters host cell metabolism and composition in other measurable ways. Cellular changes such as nucleoid degradation, cessation of net RNA synthesis and lipid remodelling are best understood in Escherichia coli phage infection systems39,40, but the extent to which these changes are shared across diverse virus–host systems is unknown. Among marine systems, cellular changes during infection have been best documented in the haptophyte alga Emiliania huxleyi. These changes include a shift from carbon fixation to the pentose phosphate pathway for viral nucleic acid synthesis30,41, extensive lipid remodelling30,42,43 and alteration of transparent exopolymer particle (TEP) production44,45. Similarly, infection has been shown to alter the cellular redox state in the marine cyanobacterium Prochlorococcus marinus strain MED4, as metabolism shifts from carbon fixation to the pentose phosphate pathway to supply nucleotides for phage replication46. Metabolite profiling has begun to reveal other global changes in cellular metabolism during infection, such as an overall increase in metabolic activity in Sulfitobacter spp., along with a stoichiometric shift to higher cellular nitrogen47. From these studies, it is clear that lytic infection drastically alters the cellular and metabolic landscape, but more work is needed to determine whether these findings apply to diverse taxa and in relevant environmental conditions. Quantifying how infected cells differ from their uninfected counterparts (for example, in cellular stoichiometry, macromolecular composition and metabolic fluxes) is not only an essential step towards integrating viruses into ecosystem models, but may also lead to new diagnostic biomarkers for measuring infection rates in natural ecosystems. Examples include the distinct lipid signatures48,49 and intracellular redox conditions50 of infected E. huxleyi cells. Being able to detect and quantify infected cells and their associated metabolic changes is crucial for estimating the pre- and post-lysis impacts of viral infection in aquatic systems.
To date, these features of viral reprogramming have largely been assessed using individual virus–host pairs under laboratory conditions. There is growing evidence that infection outcomes are highly specific to the selected virus–host pair. This specificity is evident in host transcriptional responses to infection, which, in contrast to the relatively fixed viral gene expression programme, appear to be highly variable. The same host strain of marine C. baltica shows distinct responses to different viruses27,28, and, conversely, the same virus elicits distinct responses in different host strains of both C. baltica27,28 and Synechococcus25. In Synechococcus sp. strain WH8102, infection induces expression of both putative host defence genes and genes that the virus may exploit to enhance its replication51. Moreover, several host proteins that are synthesized during infection in this same Synechococcus strain are also encoded in some phage genomes, suggesting that their expression enhances viral fitness52. Host defences against viral infection, and viral counter-defences, are diverse and largely uncharacterized53,54. Together, these findings suggest that the outcome of infection may hinge on whether the virus can repress or evade host defences, and potentially exploit host gene expression and metabolism55, and thereby efficiently produce progeny27,28. Given this specificity, a major challenge is to reconcile laboratory findings with ecological reality, in terms of the diversity and abundance of coexisting hosts and viruses.
Metabolic constraints on viral infection
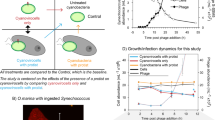
In natural environments, microorganisms are often energy or nutrient limited, based on suboptimal growth rates measured in situ56,57. This pre-infection physiological state of the host cell has long been known to affect phage production, as documented by studies linking the host growth rate and burst size in E. coli58,59,60. Beyond E. coli, a multiplicity of relationships between host physiology, environmental conditions and viral productivity have been observed in aquatic bacterial and eukaryotic algal hosts (Fig. 2). Although the data remain sparse and the findings are challenging to compare between studies and organisms (Box 2), it is clear that viral production can be slowed and/or reduced in light-limited cyanobacteria Prochlorococcus24, freshwater chlorophyte algae61 and both prasinophyte and haptophyte marine algae41,62,63,64,65,66; at suboptimal temperatures in species of the prasinophyte Micromonas62,67,68 and the haptophyte Emiliania69; and under certain nutrient (for example, nitrogen, phosphorus and iron) limitation conditions for marine bacteria Pseudoalteromonas spp.70, marine cyanobacteria Synechococcus71 and multiple Phaeocystis (haptophyte) and Micromonas strains31,63,66,72,73. By contrast, silicon stress had no detectable effect on viral burst size, but was found to accelerate virus-induced mortality in cultures of the diatom Chaetoceros tenuissimus and in natural diatom populations off the California US coast74. Extrapolating these trends more broadly remains a challenge in part because the organisms that have been studied represent an unsystematic selection of a few taxa and the conditions studied (including virus and host densities) are not necessarily reflective of those found in nature. The nature of the relationship between the host growth rate or environmental driver and viral production (that is, whether it is linear, monotonic and so forth) also remains unclear because experiments to date often compare only two conditions (for example, nutrient replete versus nutrient deplete). Furthermore, the mechanisms underlying these observed relationships between viral production and the host physiology or environment are poorly understood. It is possible that modulating the host growth rate through different forcing factors (for example, temperature, light availability or nutrient supply) leads to distinct cellular states in terms of biosynthetic machinery, resource allocations and stoichiometry, and may therefore give rise to distinct patterns of viral production. In the following sections we explore the effects of various physiological stressors on host physiology in more detail and, in turn, viral strategies to overcome host constraints, with the goal of elucidating principles that can be applied in biogeochemical models.
Simplified representation of relationships between viral productivity and host physiology or environment, shown in terms of the viral production rate (quantified as accumulation of intracellular genome copies over time24, linear regression of extracellular virions over time60,62 or virion growth rate68) (part a), viral burst size (that is, normalized yield; see Box 2 for discussion of quantification methods) (part b) and viral latent period (for example, the time period to extracellular release of new virions) (part c). The host growth rate and cellular machinery (that is, ribosomes and enzymes) can be manipulated by environmental variables (for example, temperature, light intensity or nutrient availability). Nutrient availability has the potential to alter viral production directly through limitation of substrates needed to build progeny virions or indirectly through the host growth rate, which in turn affects production yields or rates, respectively. Linear relationships are shown for simplicity but are intended to represent only the direction of correlation between viral productivity and host physiology. The slope and linearity of the relationship varies in species-specific and environment-specific ways. Interactive effects between environmental variables (for example, light and nutrient availability) can further modify the shape of the relationship. The viral burst size and latent period may be fixed traits or insensitive to the specific variables tested, as shown by the horizontal dashed grey lines (parts b and c). Data supporting these relationships were obtained from empirical studies across marine and non-marine virus–host model systems: host growth rate24,62,68, cellular machinery and dilution rate60,70 (part a); nutrient availability66,72,73,74,127,215,216, light intensity61,62,64,66,78 and temperature67 (part b); and host growth rate68, nutrient availability66,72,73,216, light intensity61,62,64,66,78,123 and temperature67 (part c).
Light and energy availability
Photosynthetic microorganisms are commonly studied experimental systems for virus–host interactions because they form the base of the food web, and their energy metabolism can be manipulated instantaneously by turning out the lights or adding specific inhibitors of photosynthesis. Among cyanobacteria, the production of cyanophages has long been known to depend on photosynthesis to varying degrees in both freshwater75 and marine76,77,78 taxa. Infected cyanobacterial cells direct energy and reducing power away from CO2 fixation and towards viral protein and DNA synthesis, with CO2 fixation ceasing well before lysis in some79,80, although not all81, freshwater cyanobacteria, and in marine Synechococcus sp. WH7803 (ref.82).
For some cyanophages, reliance on the pre-existing host energy-generation machinery is insufficient to meet their demands, either because the machinery is not abundant or efficient enough to supply viral energy demand, or because it turns over on short timescales relative to the length of the infection cycle and must be replaced to maintain activity. For these viruses, carrying and expressing their own auxiliary metabolic genes (AMGs) related to energy metabolism can offer an advantage by providing ATP and/or reducing power for viral protein synthesis, which represents the greatest energy demand for relatively small viruses including cyanophages78,83. In addition to core reaction centre proteins84,85,86,87, numerous other components of photosynthetic electron transport, along with proteins that are thought to stabilize the reaction centres, are also encoded in cyanophage genomes88 (Fig. 1b). Cyanophages also appear to contribute to phycobilin pigment biosynthesis, by encoding haeme oxygenase and bilin reductases89,90,91 that may enhance light harvesting during infection. Genes that encode core reaction centre proteins are among the best-studied viral AMGs: they are expressed at both the transcript and protein levels76,84,85,92, they affect photosystem operation and/or electron flow78,85 and they are predicted to improve cyanophage fitness93,94. To date, no single cyanophage has all of these proteins; rather, individual phages likely maintain the genes that are most crucial for viral fitness in the context of their particular host and environment.
Cyanophages also influence phototroph energy and carbon metabolism beyond the photosystems, particularly by redirecting metabolic flows through the interlocking Calvin cycle and pentose phosphate pathways (Fig. 1b). These two pathways share several bidirectional enzymes and can be viewed as running in opposition to one another, with the Calvin cycle trading reducing power (generated by photosynthesis) for fixed carbon and the pentose phosphate pathway doing the reverse. Many cyanophages encode genes for pentose phosphate pathway enzymes, including glucose 6-phosphate dehydrogenase (zwf), 6-phosphogluconate dehydrogenase (gnd) and transaldolase (tal), as well as cp12, an allosteric repressor of two Calvin cycle enzymes95,96. Expression of these genes during infection of either Prochlorococcus or Synechococcus hosts appears to promote metabolic flux through the pentose phosphate pathway at the expense of the Calvin cycle, potentially to enhance dNTP synthesis for phage replication46. Consistent with this observation, the suppression of CO2 fixation during infection was more rapid and severe for a Synechococcus-infecting cyanophage carrying the pentose phosphate pathway and cp12 genes than for one without these genes82.
Analogously, there is some evidence for continuation of host photosynthesis and reduction or cessation of CO2 fixation in diverse eukaryotic algal–virus systems. Chloroplasts (where photosynthesis is localized in algal cells) are preserved but show altered photosynthetic properties during viral infection of the prasinophyte Micromonas pusilla65 and the stramenopile Aureococcus anophagefferens32,97. As in some cyanobacteria, CO2 fixation declines early in infection in freshwater chlorophyte Chlorella variabilis61 and marine M. pusilla98. Likewise, protein synthesis of CO2-fixation machinery decreased in infected populations of E. huxleyi, but light-driven photosynthetic reactions were maintained30,41. Concurrently, expression and activity of the pentose phosphate pathway increased in a light-dependent manner, reflecting a shift towards viral nucleotide biosynthesis, in which the proportion of recycled versus de novo synthesized nucleotides was also controlled by light intensity30,41. None of the sequenced algal viruses to date encode photosynthetic reaction centre proteins, although many fewer algal viruses have had their genomes sequenced (examples given in refs16,17,18,99). We posit that one reason why these algal viruses lack photosynthesis genes is the requirement for the correct transit peptide for proteins that must be imported into the plastid, which increases the barrier to successful protein hijacking by the virus.
There are further hints that viruses manipulate energy metabolism, beyond oxygenic photosynthesis itself, from genomes of marine eukaryotic viruses. The so-called ‘giant viruses’ (>300 kb genome size) appear to encode metabolic functions that were formerly considered unique to cellular life100,101. Sugar metabolism and fermentation genes were identified in a giant virus of cosmopolitan green alga Tetraselmis spp.102. Additionally, putative microbial rhodopsin genes have been identified in the marine haptophyte Phaeocystis globosa virus PgV and in two putative eukaryotic algal virus metagenomic contigs for which the host is as yet unknown103. Rhodopsin and pigment biosynthesis genes were also found in an uncultivated virus of an uncultured choanoflagellate (predatory protist) by targeted metagenomics23. Using metagenomics, this study showed that rhodopsins are common components of giant virus genomes and demonstrated that the most common type has proton pumping activity when expressed in E. coli23. Further, a newly described family of heliorhodopsins, distantly related to known microbial rhodopsins, has also been identified in the Tara Oceans marine viral metagenomes104. Presumably, once expressed by the host, the viral rhodopsins function either to establish transmembrane proton gradients for ATP production (that is, energy conversion) or as light sensors, analogous to their microbial counterparts105,106.
Viruses that infect non-phototrophic, chemoautotrophic bacteria and archaea also appear to manipulate host energy metabolism, but few have been experimentally characterized. Metagenomic analyses have revealed that viruses infecting SUP05 bacteria, a lineage of sulfur oxidizing and denitrifying Gammaproteobacteria common throughout the deep ocean107, encode dsrC, a sulfur redox metabolism enzyme, in habitats that include hydrothermal vents and oxygen minimum zones108,109,110. Likewise, metagenomic evidence suggests that some viruses infecting another abundant marine chemoautotrophic group, the ammonia-oxidizing Thaumarchaeota, carry archaeal-like amoC genes that encode a subunit of the ammonia-oxidation machinery110,111. Recently, viruses that infect thaumarchaeon Nitrosopumilus spp. have been isolated, and although they lack amoC genes, they allow host ammonia oxidation to continue for several days post infection, which may enhance their productivity15. In all of these cases, the impact of continued host energy metabolism on viral productivity remains to be quantified.
Light can also affect viral dynamics beyond the host cell’s energy budget (reviewed in refs112,113), adding another layer of complexity to consider for time- and space-resolved models of viral biogeochemistry. For example, numerous cyanophages114,115,116 and a virus of the haptophyte alga E. huxleyi41 show reduced adsorption rates in the dark. Many phototrophs have circadian cell cycles where the time of day influences pools of nucleotides and other resources that could impact viral production117. Additionally, high light can reduce viral production in algae through cellular damage caused by reactive oxygen species41,118, but the relationship between reactive oxygen species and viral production is complex50. Light also drives the decay of free viral particles in aquatic systems119. Thus, together with its influence on adsorption and viral production, the impact of light extends to non-photosynthetic microorganisms and likely contributes to the distinct depth distributions observed for viral populations in the oceans120. Light may also have a key role in governing daily rhythms in viral infection, which have recently been documented both in freshwater121 and in coastal and open ocean37,122,123 systems, reflecting the integrated effects of light on viral infection through both phototrophic energy metabolism and photosynthesis-independent mechanisms.
Macronutrient availability
Substrate limitation has the potential to alter viral production in both growth rate-dependent mechanisms (for example, number of ribosomes, and pool sizes of precursors and enzymes)124,125 and growth rate-independent mechanisms (for example, direct substrate availability) (Fig. 2), depending on the needs of the host and the virus. Few empirical measurements of viral elemental composition have been made, and these values are likely to vary with virion size, morphology and the presence of a lipid envelope. Predictions of stoichiometry for phage126 and algal virus127 particles suggest enrichment in nitrogen and phosphorus compared with host cells, implying that viruses must concentrate these elements to reproduce. But to what extent are viruses restricted to recycling intracellular nucleotides and amino acids, compared with de novo synthesis using newly acquired nutrients? Across virus–phytoplankton systems, the host genome size is a strong predictor of viral burst size128,129, suggesting that viral production depends, in part, on intracellular nucleotide pools. Consistent with this, radiotracer experiments with several marine phage–host pairs suggested that the phosphorus in phage DNA came mostly from host nucleotides130. If viruses are limited by intracellular phosphorus availability, then it should be possible to modulate the burst size by changing cellular carbon:phosphorus ratios independent of the growth rate — indeed, this was observed in a freshwater virus–alga system127. Thus, at least in some systems, host resource pools have the potential to constrain viral productivity.
At the same time, there is also evidence that some marine viruses can take advantage of resources outside the host cell. Early studies of E. coli phages demonstrated that viral replication requires extracellular nitrogen and phosphorus58, even when the hosts are not starved, as the bulk of coliphage DNA nitrogen is extracellular in origin131. Recently, isotopic labelling experiments in the haptophyte E. huxleyi and marine Synechococcus revealed a similar reliance on extracellular nutrients by tracing the incorporation of carbon and nitrogen into viral particles132 or of nitrogen into specific viral proteins52. Although data are limited, for phosphorus133 and nitrogen52, phages infecting E. coli and Synechococcus, respectively, appear to source relatively more nutrients from the host cell biomass during the early stages of infection, and shift to extracellularly derived nutrients as the infection proceeds. Which pools of host biomolecules are accessible to the virus, which host biomolecules are potentially off-limits (for example, ribosomes) and how this accessibility is controlled are all unknown. Ultimately, the overall balance between intracellular and extracellular resources for viral replication likely varies with environmental nutrient availability as well as with the host growth rate and physiological state at the time of infection.
Viral exploitation of extracellular resources, presumably to alleviate a resource bottleneck during infection, is also reflected in the variety of AMGs related to nutrient acquisition. Many marine cyanophage isolates encode transport machinery for phosphorus, in particular the substrate binding protein PstS and, less commonly, a protein bearing similarity to a putative alkaline phosphatase134. Cyanophage phosphorus-uptake genes are upregulated during infection of phosphorus-starved host cells33 by the host PhoBR two-component system135, which activates the expression of genes for phosphorus acquisition and metabolism in response to phosphorus limitation136. Hence, the phage is able to recognize and specifically respond to phosphorus limitation inside the host cell. Cyanophage-encoded phosphorus-acquisition genes tend to occur more frequently in viral genomes from phosphorus-limited regions of the oceans134,137. In these regions, cyanobacterial host cells likely have reduced intracellular phosphorus content138,139, presumably forcing phages to rely more heavily on extracellular uptake to replicate. Similarly, a haptophyte algal virus isolated from a bloom of E. huxleyi in the English Channel encodes a phosphate permease that is absent in a related strain isolated from a Norwegian fjord99,140, and this gene may help facilitate infection in phosphorus-limited waters.
Transporters for both phosphorus and nitrogen are also present in a subset of viruses that infect prasinophyte algae, interestingly in hosts that generally come from nutrient-replete environments141,142. Although there is much unexplored eukaryotic viral diversity in the oceans, the only nitrogen transporter identified among all sequenced viral genomes to date is encoded and expressed by the virus OtV6, which infects the picoeukaryotic prasinophyte alga Ostreococcus tauri141. Surprisingly, both the host and the virus were isolated from an extremely nitrogen-rich environment, an oyster lagoon in coastal France. The viral protein is related to a broad family of ammonium transporters common to all eukaryotic life, and, in this case, appears to have been acquired directly from the algal host. Functional characterization indicates that the protein broadens the diversity of nitrogen sources that can be accessed by the host and increases substrate affinity over the host transporter141.
By contrast, AMGs encoding nitrogen transport proteins are absent in phages, at least to date. This absence, along with evidence that phages acquire substantial extracellular nitrogen, suggests that the existing host cell nitrogen uptake machinery and recycling of intracellular stores are generally sufficient to meet the demands of viral production. One major pool of nitrogen that cyanophage may be able to access is phycobilisomes, which can account for up to 50% of the soluble protein in a cyanobacterial cell143. Proteins involved in phycobilisome degradation have been identified in several freshwater cyanophage genomes and marine metagenomes, and in some cases biochemically validated144,145,146, and their degradation products have been observed in viral lysates147. These proteins may help supply nitrogen for viral production, although how viruses balance their need for substrates with their need for continued host photosynthesis and metabolism is unknown.
Micronutrient availability
Besides nitrogen and phosphorus, viruses also appear to influence cellular pathways related to the synthesis of cofactors and other small molecules, highlighting potential bottlenecks in viral production. One such molecule is cobalamin (vitamin B12)148, a cofactor used in the enzymatic reduction of ribonucleotides to deoxyribonucleotides for viral replication. Many marine bacteria produce vitamin B12 de novo, whereas eukaryotic phytoplankton do not, making the exchange of these compounds an ecologically important cross-kingdom microbial interaction149,150. Several cyanophages and an archaeal virus encode a putative cobS gene134,151, the product of which is predicted to catalyse the last step of vitamin B12 synthesis in bacteria. However, the cyanophage cobS is phylogenetically distinct from the host gene, suggesting that it was not acquired directly from its host and that it may have different functional properties152. As with most AMGs described above, the viral cobS protein product has not been biochemically characterized and hence its role in vitamin B12 cycling and viral production remains speculative.
Trace metals, in particular iron, are known to limit phytoplankton growth in much of the global ocean153. Viral particles are not known to directly incorporate trace metals and other micronutrients, although it has been proposed that viral particles can act as important metal ligands154. Trace metals may impact viral production indirectly by controlling the overall host growth rate or the activity of specific cofactor-requiring enzymes. Iron limitation has been shown to reduce the burst size in algal P. globosa and Micromonas species73. However, high trace metal concentrations can also be toxic and inhibit virus replication, as seen for copper and viruses infecting the marine haptophyte E. huxleyi155. Thus, trace metals undoubtedly impact host physiology and thereby viral infection dynamics, but the molecular mechanisms and ecosystem-level consequences of trace metal–virus interactions are unknown.
Functional divergence in host and viral metabolism
As our catalogue of putative AMGs continues to grow, there is greater need for biochemical characterization to assess viral contributions to the metabolism of infected cells and, by extension, to biogeochemistry. Even for homologous proteins shared by the host and the virus, the version that maximizes viral fitness may not necessarily maximize host fitness, and vice versa, given the different biochemical environments of infected and uninfected cells, and distinct constraints acting on cells and viruses. Hence, homologues encoded by the host and the virus may have very different substrate specificities, kinetics or other properties. Indeed, AMGs characterized to date are often divergent from their host homologues. Examples from cyanophages include phycoerythrobilin synthase (PebS), which combines the activities of two host enzymes89; transaldolase (TalC), which is both much shorter in length and less efficient than the host version46; and phycobiliprotein lyase (CpeT), which does not appear to catalyse the same reaction as the host version156. Examples from algal viruses include the viral-encoded ammonium transporter discussed above141 and viral serine palmitoyltransferase (vSPT), an enzyme involved in sphingolipid biosynthesis that differs in substrate specificity from the host version and thereby alters the chemical nature of the infected cell42. At an even finer scale, biochemical diversity of vSPT enzymes is evident among viruses that infect a common E. huxleyi host, and these biochemical differences can, in turn, influence competitive interactions among viral strains157. Functional characterization of viral AMGs, through enzymological46, physiological42 and genetic approaches, is essential for understanding how infection alters cellular metabolism, and may point towards new molecular diagnostics of infection that can be applied in natural ecosystems48.
Viral impacts on biogeochemical cycles
The biogeochemical influence of viruses begins at the moment of infection, due to metabolic remodelling of the host cell8, and continues even after cellular lysis, as the viral progeny and cellular debris disperse into the surrounding environment where they become food for the wider microbial community, catalyse biogeochemical transformations and initiate new infections4,158,159. Competition for resources among the members of this microbial community, in turn, determines the nutrient status and physiological state of the next host cell that viral progeny will ultimately infect. Numerous field studies, primarily in aquatic ecosystems, have documented nutrient stimulation of primary and/or bacterial productivity that is accompanied by increases in viral productivity or abundance160,161,162. It is becoming increasingly clear that viral reproduction in microbial ecosystems is both influenced by and is a contributor to biogeochemical processes at spatial and temporal scales well beyond the infection of individual host cells.
Nutrient recycling versus export
A prevailing concept in considerations of the biogeochemical impact of marine viruses has been the ‘viral shunt’4,5. This hypothesis emphasizes that lytic infections return lysed host cell biomass back to the dissolved phase, as dissolved organic matter (DOM), which in turn regenerates nutrients for microorganisms and ‘shunts’ them away from grazers and higher trophic levels (Fig. 3). Although measuring the strength or large-scale impact of the viral shunt is challenging, modelling suggests that it can increase overall productivity of marine ecosystems by enhancing the efficiency of nutrient recycling that enables the majority of marine primary productivity163, a feature that might be particularly important during nutrient-stimulated bloom events160. An ecosystem population dynamics model has recently been used to assess the viral shunt from mortality measurements in the California Current ecosystem, calculating carbon transfer from primary producers to viruses, grazers and the DOM pool164, and silicon limitation of diatom blooms in this ecosystem has been suggested to accelerate viral termination of the bloom and thereby strengthen the viral shunt74.
Schematic of two contrasting scenarios for carbon and nutrient fluxes driven by viral infection of primary producers in aquatic ecosystems. The ‘viral shunt’ (left) emphasizes release of cellular constituents to the dissolved organic matter (DOM) pool, which fuels heterotrophic microbial production and nutrient recycling through the microbial loop, at the expense of grazing and higher trophic levels. The ‘viral shuttle’ (right) describes ways in which viral activity could facilitate carbon export to the deep ocean, including direct grazing on viral particles and infected cells, as well as particle aggregation and sinking driven by the release of lysis products and/or virus-induced alterations in host physiology such as transparent exopolymeric polysaccharide (TEP) production. An oversimplification here is that grazers are also known to be infected by viruses (not depicted). Orange arrows indicate uptake of macronutrients (nitrogen and phosphorus), and their preferential incorporation into protein-rich and nucleic-acid-rich viral particles, leaving relatively nitrogen-depleted and phosphorus-depleted byproducts of infection and lysis (dashed arrows).
Owing to extensive biochemical remodelling during infection, DOM released by the infected cell or by viral lysis appears compositionally distinct compared with that produced through prey cell breakage during grazing (‘sloppy feeding’) or exudation by growing phytoplankton47,147,165. Although data are limited, it appears that cell wall degradation and stimulated nucleotide synthesis47, exopolysaccharide degradation166, lipid remodelling42,48 and specific proteolysis of abundant protein complexes147 contribute to these distinct DOM signatures. Relative enrichments of amino acids47,167 and proteinaceous material147,165,168 have been observed in multiple types of viral lysate, suggesting that viral lysis may be a particularly important mechanism for nitrogen recycling through the viral shunt169. Similarly, viral lysis has been shown to release iron in highly bioavailable forms which can be taken up by other microbial cells170,171, and organic phosphorus released from viral lysis has been shown to support the phosphorus demand of uninfected but phosphorus-limited cultures of marine Vibrio spp.172. In addition to these labile components, viral lysis is thought to yield long-lived, recalcitrant DOM as well173. To date, most studies of viral effects on DOM have been performed in nutrient-replete laboratory cultures; how these DOM signatures covary with cellular composition, for example, as elemental stoichiometry shifts across resource gradients174, remains to be tested. More recently, evidence has emerged that viral infection may actually enhance, rather than reduce, carbon export from surface to deep waters, prompting the notion of a ‘viral shuttle’175,176 (Fig. 3). This viral shuttling might happen either because infected cells and/or lysis products aggregate and sink at higher rates, or due to enhanced grazing on viral particles and/or infected cells. Virus-induced production of aggregates is attributed to proteinaceous material in the diatom C. tenuissimus168. In other hosts, viral infection stimulates production of TEPs — the ‘marine snow’ known to act as a glue for particulate matter and to enhance aggregation into larger and denser particles45,165. Field observations of several mesoscale (~100 km) E. huxleyi blooms in the North Atlantic showed that TEP concentrations increased during the early stages of infection, and infected cells were preferentially entrained in sinking aggregates; these aggregates were ballasted by the extracellular calcium carbonate scales characteristic of this host44,177. These sinking particles shuttle nutrients out of the surface ocean, which potentially makes them available to larger grazers in the mesopelagic and/or shifts viral lysis and shunt-style nutrient recycling deeper into the water column. The calcified scales covering E. huxleyi cells appear to protect some strains from viral infection and scale fragments act as effective adsorbers of viral particles; the viruses, in turn, are able to induce as-yet-unidentified infochemicals that reduce host calcification and render them more susceptible to infection178, suggesting that export through the viral shuttle can depend on the complex interplay between viral infection and carbonate ballast production.
There is also evidence that two major death processes for aquatic microorganisms — viral infection and predator grazing — can interact in unexpected ways. Virus-infected E. huxleyi cells were found to be grazed at higher rates than uninfected cells by the dinoflagellate Oxyrrhis marina179, but at lower rates by the copepod Acartia tonsa180. Copepods ingested infected E. huxleyi cells at high rates under both laboratory conditions and during a bloom event in the North Atlantic, which could act as a vector for viral dispersal181. Marine viruses themselves are actively ingested by multiple types of predators181,182,183, indicating that viral particles can have a role in the ‘classical’ marine food web. The rate of removal of virus particles by aquatic protists appears to depend on the specific virus and host strains183, as well as the feeding mode of the protist158. Taken altogether, it seems that the viral shunt and the viral shuttle likely operate simultaneously in many sunlit aquatic ecosystems, their relative importance waxing and waning depending on host traits (for example, cell size or the presence of a ballast), viral effects on metabolism (for example, TEP production) and environmental conditions. Diagnostic biomolecular tools for some aspects of these processes are emerging48,177 and will provide novel and needed constraints on the rates and variability of viral biogeochemistry.
Viruses in global biogeochemical models
At this point, we have outlined mechanistically and conceptually how viruses potentially alter biogeochemistry through metabolic reprogramming and lysis. At the cellular level, metabolically detailed models of infection physiology have been constructed for a few virus–host model systems40,41 and could be expanded to others. However, virus impacts on ecosystem-level processes are just beginning to be incorporated into biogeochemical models163,164,184,185. A major caveat is that models of viral infection strategies and replication dynamics in natural communities are often based on laboratory studies of select virus–host model systems, in conditions that are physically, chemically and biologically unrealistic. Because our experimental systems are so limited, it has been challenging to extract common principles of infection that can serve as a foundation for improved ecosystem models. Accordingly, none of the models used to predict global responses to CO2 inputs for the Intergovernmental Panel on Climate Change Assessment Report186 include an explicit representation of viruses187. Instead, some models implicitly represent viral activity, for instance, within the conversion of particulate nutrients to dissolved nutrients153,188,189. Yet, given the multifarious ways in which changes in viral activity could either amplify or dampen climate forcing — including influencing the biological carbon pump in the ocean or the production of marine aerosols and reactive trace gases190, along with the temperature sensitivity of virus–host interactions68,69 — a mechanistic, quantitative framework for including viruses in ecosystem and large-scale biogeochemical models is needed. Developing such a framework requires substantial knowledge from both laboratory and field studies, which to date exists for few virus–host systems157.
Treatment of infected host cells as a separate functional class in biogeochemical models may not be necessary for all purposes, but with the expanding appreciation of how extensive and prolonged remodelling during infection can be, its integration into large-scale productivity models seems warranted. For instance, the global impact of cessation of CO2 fixation during phage infection of marine cyanobacteria could amount to several petagrams of carbon per year82, although this estimate does not account for indirect stimulatory effects of infection on productivity163. One challenge to achieving this effectively may be the fact that physiologically distinct infection strategies can be pursued by closely-related viruses infecting the same host45,191,192, so there may be multiple, metabolically distinct ‘infected host types’ per ‘uninfected host type’, which could lead to a rapid expansion of model complexity, without necessarily providing better predictive power or greater mechanistic insight. These challenges underscore the importance of elucidating the overarching principles and commonalities that govern viral metabolic remodelling during infection and its consequences for processes such as nutrient recycling, particle aggregation and grazing.
An important area of future exploration is the effect of viruses on the major element stoichiometry (that is, carbon:nitrogen:phosphorus ratios) of organic matter exported from the euphotic zone of the ocean. The export ratios of these elements determine the efficiency of the biological pump that stores vast amounts of carbon in the ocean’s deep interior. The relatively high concentration of carbon-rich exopolysaccharides in some infected hosts and viral lysates44,45,177 would be expected to enhance carbon:nitrogen or carbon:phosphorus ratios in exported organic matter resulting from viral infection; however, other observations that carbon is remineralized more quickly than nitrogen and phosphorus in lysate would suggest that viral activity actually decreases carbon export171. Although both observational and modelling studies have shown that changing the elemental stoichiometry of biological particulate material can have a major impact on global biogeochemical cycles174,193,194, whether viral infection alters the stoichiometry of export on large scales is relatively unexplored. Understanding this potential alteration will require in-depth tracing of nutrient sourcing and flows through the infection process52 as well as field measurements of virally mediated export events177 in order to quantify the extent and mechanism of viral alteration of export fluxes.
Conclusions and outlook
Viral replication involves metabolic remodelling of the infected host cell — often, although not always, to a drastic degree — and this remodelling creates a functionally new type of cell for the period of the infection7,8. Therefore, the biogeochemical impacts of viral infection are not limited to host cell killing and the release of lysis products, but begin the moment the viral genetic material enters the cell. Although we have focused on lytic infection in this Review, these impacts are likely to be very different depending on the infection strategy, and ‘alternative’ modes beyond classic lytic infection are probably widespread in nature. This raises the sobering possibility that we have yet to experimentally characterize the infection dynamics most relevant to natural systems. Nevertheless, recent studies have begun to shed light on our blindspots by exploring a broader diversity of virus–host systems, physiological states and environmental conditions, and by beginning to assess viral infection rates and impacts in the wild. Expanding these efforts is essential if we are to elucidate fundamental principles that can guide our efforts to include viral activity in global biogeochemical models. Given the power of viruses to reprogramme cells, and potentially ecosystems, integrating them into global models is an important step towards better predicting the consequences of regional- and global-scale environmental change.
References
Jacob, F. The Logic of Life: A History of Heredity (Princeton Univ. Press, 1973).
Falkowski, P. G., Fenchel, T. & DeLong, E. F. The microbial engines that drive Earth’s biogeochemical cycles. Science 320, 1034–1039 (2008).
Pomeroy, L. R. The ocean’s food web, a changing paradigm. Bioscience 24, 499–504 (1974).
Fuhrman, J. A. Marine viruses and their biogeochemical and ecological effects. Nature 399, 541–548 (1999).
Wilhelm, S. W. & Suttle, C. A. Viruses and nutrient cycles in the sea. Bioscience 49, 781–788 (1999).
Bidle, K. D. Programmed cell death in unicellular phytoplankton. Curr. Biol. 26, R594–R607 (2016).
Forterre, P. The virocell concept and environmental microbiology. ISME J. 7, 233–236 (2013).
Rosenwasser, S., Ziv, C., Creveld, S. G. van. & Vardi, A. Virocell metabolism: metabolic innovations during host–virus interactions in the ocean. Trends Microbiol. 24, 821–832 (2016).
Suttle, C. A. Marine viruses—major players in the global ecosystem. Nat. Rev. Microbiol. 5, 801–812 (2007).
Breitbart, M. et al. Genomic analysis of uncultured marine viral communities. Proc. Natl Acad. Sci. USA 99, 14250–14255 (2002).
Angly, F. E. et al. The marine viromes of four oceanic regions. PLOS Biol. 4, 2121–2131 (2006).
Krupovic, M., Cvirkaite-Krupovic, V., Iranzo, J., Prangishvili, D. & Koonin, E. V. Viruses of archaea: structural, functional, environmental and evolutionary genomics. Virus Res. 244, 181–193 (2018).
Paez-Espino, D. et al. Uncovering Earth’s virome. Nature 536, 425–430 (2016).
Gregory, A. C. et al. Marine DNA viral macro- and microdiversity from pole to pole. Cell 177, 1109–1123 (2019).
Kim, J.-G. et al. Spindle-shaped viruses infect marine ammonia-oxidizing thaumarchaea. Proc. Natl Acad. Sci. USA 116, 15645–15650 (2019). This study presents the first reported isolation of viruses infecting widespread marine archaea, demonstrating the continuation of ammonium oxidation activity during infection and a chronic infection strategy distinct from that of the lytic bacteriophage.
Derelle, E. et al. Diversity of viruses infecting the green microalga Ostreococcus lucimarinus. J. Virol. 89, 5812–5821 (2015).
Moniruzzaman, M. et al. Genome of brown tide virus (AaV), the little giant of the Megaviridae, elucidates NCLDV genome expansion and host–virus coevolution. Virology 466–467, 60–70 (2014).
Santini, S. et al. Genome of Phaeocystis globosa virus PgV-16T highlights the common ancestry of the largest known DNA viruses infecting eukaryotes. Proc. Natl Acad. Sci. USA 110, 10800–10805 (2013).
Deeg, C. M., Chow, C. E. T. & Suttle, C. A. The kinetoplastid-infecting Bodo saltans virus (Bsv), a window into the most abundant giant viruses in the sea. eLife 7, e33014 (2018).
Claverie, J.-M. & Abergel, C. Mimiviridae: an expanding family of highly diverse large dsDNA viruses infecting a wide phylogenetic range of aquatic eukaryotes. Viruses 10, 506 (2018).
Coy, S., Gann, E., Pound, H., Short, S. & Wilhelm, S. Viruses of eukaryotic algae: diversity, methods for detection, and future directions. Viruses 10, 487 (2018).
Hingamp, P. et al. Exploring nucleo-cytoplasmic large DNA viruses in Tara Oceans microbial metagenomes. ISME J. 7, 1678–1695 (2013).
Needham, D. M. et al. A distinct lineage of giant viruses brings a rhodopsin photosystem to unicellular marine predators. Proc. Natl Acad. Sci. USA 116, 20574–20583 (2019).
Lindell, D. et al. Genome-wide expression dynamics of a marine virus and host reveal features of co-evolution. Nature 449, 83–86 (2007).
Doron, S. et al. Transcriptome dynamics of a broad host-range cyanophage and its hosts. ISME J. 10, 1437–1455 (2016).
Morimoto, D., Kimura, S., Sako, Y. & Yoshida, T. Transcriptome analysis of a bloom-forming cyanobacterium Microcystis aeruginosa during Ma-LMM01 phage infection. Front. Microbiol. 9, 2 (2018).
Howard-Varona, C. et al. Regulation of infection efficiency in a globally abundant marine Bacteriodetes virus. ISME J. 11, 284–295 (2017).
Howard-Varona, C. et al. Multiple mechanisms drive phage infection efficiency in nearly identical hosts. ISME J. 12, 1605–1618 (2018).
Allen, M. J. et al. Locus-specific gene expression pattern suggests a unique propagation strategy for a giant algal virus. J. Virol. 80, 7699–7705 (2006).
Rosenwasser, S. et al. Rewiring host lipid metabolism by large viruses determines the fate of Emiliania huxleyi, a bloom-forming alga in the ocean. Plant Cell 26, 2689–2707 (2014).
Bachy, C. et al. Transcriptional responses of the marine green alga Micromonas pusilla and an infecting prasinovirus under different phosphate conditions. Environ. Microbiol. 20, 2898–2912 (2018).
Moniruzzaman, M., Gann, E. R. & Wilhelm, S. W. Infection by a giant virus (AaV) induces widespread physiological reprogramming in Aureococcus anophagefferens CCMP1984-A harmful bloom algae. Front. Microbiol. 9, 752 (2018).
Lin, X., Ding, H. & Zeng, Q. Transcriptomic response during phage infection of a marine cyanobacterium under phosphorus-limited conditions. Environ. Microbiol. 18, 450–460 (2016).
Rosenwasser, S. et al. Unmasking cellular response of a bloom-forming alga to viral infection by resolving expression profiles at a single-cell level. PLOS Pathog. 15, e1007708 (2019).
Sieradzki, E. T., Ignacio-Espinoza, J. C., Needham, D. M., Fichot, E. B. & Fuhrman, J. A. Dynamic marine viral infections and major contribution to photosynthetic processes shown by regional and seasonal picoplankton metatranscriptomes. Nat. Commun. 10, 1169 (2019).
Moniruzzaman, M. et al. Virus–host relationships of marine single-celled eukaryotes resolved from metatranscriptomics. Nat. Commun. 8, 16054 (2017).
Aylward, F. O. et al. Diel cycling and long-term persistence of viruses in the ocean’s euphotic zone. Proc. Natl Acad. Sci. USA 114, 11446–11451 (2017).
Kolody, B. C. et al. Diel transcriptional response of a California Current plankton microbiome to light, low iron, and enduring viral infection. ISME J. 13, 2817–2833 (2019).
Kutter, E. et al. From host to phage metabolism: hot tales of phage T4’s takeover of E. coli. Viruses 10, 387 (2018).
Yin, J. & Redovich, J. Kinetic modeling of virus growth in cells. Microbiol. Mol. Biol. Rev. 82, e00066-17 (2018).
Thamatrakoln, K. et al. Light regulation of coccolithophore host–virus interactions. New Phytol. 221, 1289–1302 (2019). Based on photophysiology and biochemical measurements during E. huxleyi viral infection, this study suggests that viral replication is controlled by a light-dependent trade-off between host nucleotide recycling and de novo synthesis.
Ziv, C. et al. Viral serine palmitoyltransferase induces metabolic switch in sphingolipid biosynthesis and is required for infection of a marine alga. Proc. Natl Acad. Sci. USA 113, E1907–E1916 (2016).
Malitsky, S. et al. Viral infection of the marine alga Emiliania huxleyi triggers lipidome remodeling and induces the production of highly saturated triacylglycerol. New Phytol. 210, 88–96 (2016).
Vardi, A. et al. Host–virus dynamics and subcellular controls of cell fate in a natural coccolithophore population. Proc. Natl Acad. Sci. USA 109, 19327–19332 (2012).
Nissimov, J. I. et al. Dynamics of transparent exopolymer particle (TEP) production and aggregation during viral infection of the coccolithophore, Emiliania huxleyi. Environ. Microbiol. 20, 2880–2897 (2018).
Thompson, L. R. et al. Phage auxiliary metabolic genes and the redirection of cyanobacterial host carbon metabolism. Proc. Natl Acad. Sci. USA 108, E757–E764 (2011). This paper shows that cyanophages encode a Calvin cycle inhibitor and transaldolase with enzymological properties different from their host homologues, demonstrating the importance of the pentose phosphate pathway during infection.
Ankrah, N. Y. D. et al. Phage infection of an environmentally relevant marine bacterium alters host metabolism and lysate composition. ISME J. 8, 1089–1100 (2014). This paper uses metabolomics to quantify redirection of metabolic fluxes during phage infection of a marine α-proteobacterium, and consequent compositional alteration of dissolved material released by lysis.
Schleyer, G. et al. In plaque-mass spectrometry imaging of a bloom-forming alga during viral infection reveals a metabolic shift towards odd-chain fatty acid lipids. Nat. Microbiol. 4, 527–538 (2019).
Hunter, J. E., Frada, M. J., Fredricks, H. F., Vardi, A. & Van Mooy, B. A. S. Targeted and untargeted lipidomics of Emiliania huxleyi viral infection and life cycle phases highlights molecular biomarkers of infection, susceptibility, and ploidy. Front. Mar. Sci. 2, 81 (2015).
Schieler, B. M. et al. Nitric oxide production and antioxidant function during viral infection of the coccolithophore Emiliania huxleyi. ISME J. 13, 1019–1031 (2019).
Fedida, A. & Lindell, D. Two Synechococcus genes, two different effects on cyanophage infection. Viruses 9, 136 (2017).
Waldbauer, J. R. et al. Nitrogen sourcing during viral infection of marine cyanobacteria. Proc. Natl Acad. Sci. USA 116, 15590–15595 (2019). This proteomics study quantitatively tracks nitrogen incorporation during phage infection of Synechococcus, showing that substantial amounts of phage protein nitrogen are acquired from the environment after infection begins and incorporated via de novo amino acid synthesis.
Doron, S. et al. Systematic discovery of antiphage defense systems in the microbial pangenome. Science 359, eaar4120 (2018).
Koonin, E. V., Makarova, K. S. & Wolf, Y. I. Evolutionary genomics of defense systems in Archaea and Bacteria. Annu. Rev. Microbiol. 71, 233–261 (2017).
Schatz, D. et al. Hijacking of an autophagy-like process is critical for the life cycle of a DNA virus infecting oceanic algal blooms. New Phytol. 204, 854–863 (2014).
Hoehler, T. M. & Jørgensen, B. B. Microbial life under extreme energy limitation. Nat. Rev. Microbiol. 11, 83–94 (2013).
Kirchman, D. L. Processes in Microbial Ecology 99–116 (Oxford Univ. Press, 2012).
Cohen, S. S. Growth requirements of bacterial viruses. Bacteriol. Rev. 13, 1–24 (1949).
Hadas, H., Einav, M., Fishov, I. & Zaritsky, A. Bacteriophage T4 development depends on the physiology of its host Escherichia coli. Microbiology 143, 179–185 (1997).
You, L., Suthers, P. F. & Yin, J. Effects of Escherichia coli physiology on growth of phage T7 in vivo and in silico. J. Bacteriol. 184, 1888–1894 (2002).
Van Etten, J. L., Burbank, D. E., Xia, Y. & Meints, R. H. Growth cycle of a virus, PBCV-1, that infects Chlorella-like algae. Virology 126, 117–125 (1983).
Piedade, G. J., Wesdorp, E. M., Montenegro-Borbolla, E., Maat, D. S. & Brussaard, C. P. D. Influence of irradiance and temperature on the virus MpoV-45T infecting the arctic picophytoplankter Micromonas polaris. Viruses 10, 676 (2018).
Bratbak, G., Jacobsen, A., Heldal, M., Nagasaki, K. & Thingstad, T. F. Virus production in Phaeocystis pouchetii and its relation to host cell growth and nutrition. Aquat. Microb. Ecol. 16, 1–9 (1998).
Baudoux, A.-C. & Brussaard, C. P. D. Influence of irradiance on virus–algal host interactions. J. Phycol. 44, 902–908 (2008).
Brown, C. M., Campbell, D. A. & Lawrence, J. E. Resource dynamics during infection of Micromonas pusilla by virus MpV-Sp1. Environ. Microbiol. 9, 2720–2727 (2007).
Maat, D. S., de Blok, R. & Brussaard, C. P. D. Combined phosphorus limitation and light stress prevent viral proliferation in the phytoplankton species Phaeocystis globosa, but not in Micromonas pusilla. Front. Mar. Sci. 3, 160 (2016).
Maat, D. S. et al. Characterization and temperature dependence of arctic Micromonas polaris viruses. Viruses 9, 134 (2017).
Demory, D. et al. Temperature is a key factor in Micromonas–virus interactions. ISME J. 11, 601–612 (2017).
Kendrick, B. J. et al. Temperature-induced viral resistance in Emiliania huxleyi (Prymnesiophyceae). PLOS ONE 9, e112134 (2014).
Middelboe, M. Bacterial growth rate and marine virus–host dynamics. Microb. Ecol. 40, 114–124 (2000).
Wilson, W. H., Carr, N. G. & Mann, N. H. The effect of phosphate status on the kinetics of cyanophage infection in the oceanic cyanobacterium Synechococcus sp. WH7803. J. Phycol. 32, 506–516 (1996).
Maat, D. S. & Brussaard, C. P. D. Both phosphorus and nitrogen limitation constrain viral proliferation in marine phytoplankton. Aquat. Microb. Ecol. 77, 87–97 (2016).
Slagter, H. A., Gerringa, L. J. A. & Brussaard, C. P. D. Phytoplankton virus production negatively affected by iron limitation. Front. Mar. Sci. 3, 156 (2016).
Kranzler, C. F. et al. Silicon limitation facilitates virus infection and mortality of marine diatoms. Nat. Microbiol. https://doi.org/10.1038/s41564-019-0502-x (2019). Using both cultured isolates and field observations, this study shows that silicon stress can accelerate virus-induced mortality of marine diatoms, potentially promoting nutrient recycling via the viral shunt.
Padan, E. & Shilo, M. Cyanophages–viruses attacking blue–green algae. Bacteriol. Rev. 37, 343–370 (1973).
Lindell, D., Jaffe, J. D., Johnson, Z. I., Church, G. M. & Chisholm, S. W. Photosynthesis genes in marine viruses yield proteins during host infection. Nature 438, 86–89 (2005).
Thompson, L. R., Zeng, Q. & Chisholm, S. W. Gene expression patterns during light and dark infection of Prochlorococcus by cyanophage. PLOS ONE 11, e0165375 (2016).
Puxty, R. J., Evans, D. J., Millard, A. D. & Scanlan, D. J. Energy limitation of cyanophage development: implications for marine carbon cycling. ISME J. 12, 1273–1286 (2018). This study demonstrates that cyanophages modulate expression of photosynthesis-related accessory metabolic genes in response to light intensity, suggesting energy limitation of phage productivity and a basis for diel and seasonal patterns of virus-induced mortality.
Ginzburg, D., Padan, E. & Shilo, M. Effect of cyanophage infection on CO2 photoassimilation in Plectonema boryanum. J. Virol. 2, 695–701 (1968).
Adolph, K. W. & Haselkorn, R. Photosynthesis and the development of blue–green algal virus N-1. Virology 47, 370–374 (1972).
Mackenzie, J. J. & Haselkorn, R. Photosynthesis and the development of blue–green algal virus SM-1. Virology 49, 517–521 (1972).
Puxty, R. J., Millard, A. D., Evans, D. J. & Scanlan, D. J. Viruses inhibit CO2 fixation in the most abundant phototrophs on Earth. Curr. Biol. 26, 1585–1589 (2016).
Mahmoudabadi, G., Milo, R. & Phillips, R. The energetic cost of building a virus. Proc. Natl Acad. Sci. USA 114, E4324–E4333 (2017).
Sharon, I. et al. Photosystem I gene cassettes are present in marine virus genomes. Nature 461, 258–262 (2009).
Fridman, S. et al. A myovirus encoding both photosystem I and II proteins enhances cyclic electron flow in infected Prochlorococcus cells. Nat. Microbiol. 2, 1350–1357 (2017).
Mann, N. H. Phages of the marine cyanobacterial picophytoplankton. FEMS Microbiol. Rev. 27, 17–34 (2003).
Lindell, D. et al. Transfer of photosynthesis genes to and from Prochlorococcus viruses. Proc. Natl Acad. Sci. USA 101, 11013–11018 (2004).
Puxty, R. J., Millard, A. D., Evans, D. J. & Scanlan, D. J. Shedding new light on viral photosynthesis. Photosynth. Res. 126, 71–97 (2015).
Dammeyer, T., Bagby, S. C., Sullivan, M. B., Chisholm, S. W. & Frankenberg-Dinkel, N. Efficient phage-mediated pigment biosynthesis in oceanic cyanobacteria. Curr. Biol. 18, 442–448 (2008).
Ledermann, B., Béjà, O. & Frankenberg-Dinkel, N. New biosynthetic pathway for pink pigments from uncultured oceanic viruses. Environ. Microbiol. 18, 4337–4347 (2016).
Ledermann, B. et al. Evolution and molecular mechanism of four-electron reducing ferredoxin-dependent bilin reductases from oceanic phages. FEBS J. 285, 339–356 (2018).
Clokie, M. R. J. et al. Transcription of a ‘photosynthetic’ T4-type phage during infection of a marine cyanobacterium. Environ. Microbiol. 8, 827–835 (2006).
Hellweger, F. L. Carrying photosynthesis genes increases ecological fitness of cyanophage in silico. Environ. Microbiol. 11, 1386–1394 (2009).
Bragg, J. G. & Chisholm, S. W. Modeling the fitness consequences of a cyanophage-encoded photosynthesis gene. PLOS ONE 3, e3550 (2008).
Hurwitz, B. L. & U’Ren, J. M. Viral metabolic reprogramming in marine ecosystems. Curr. Opin. Microbiol. 31, 161–168 (2016).
Crummett, L. T., Puxty, R. J., Weihe, C., Marston, M. F. & Martiny, J. B. H. The genomic content and context of auxiliary metabolic genes in marine cyanomyoviruses. Virology 499, 219–229 (2016).
Brown, C. M. & Bidle, K. D. Attenuation of virus production at high multiplicities of infection in Aureococcus anophagefferens. Virology 466–467, 71–81 (2014).
Waters, R. E. & Chan, A. T. Micromonas pusilla virus: the virus growth cycle and associated physiological events within the host cells; host range mutation. J. Gen. Virol. 63, 199–206 (1982).
Allen, M. J., Schroeder, D. C., Donkin, A., Crawfurd, K. J. & Wilson, W. H. Genome comparison of two coccolithoviruses. Virol. J. 3, 15 (2006).
Abrahão, J. et al. Tailed giant Tupanvirus possesses the most complete translational apparatus of the known virosphere. Nat. Commun. 9, 749 (2018).
Fischer, M. G., Allen, M. J., Wilson, W. H. & Suttle, C. A. Giant virus with a remarkable complement of genes infects marine zooplankton. Proc. Natl Acad. Sci. USA 107, 19508–19513 (2010).
Schvarcz, C. R. & Steward, G. F. A giant virus infecting green algae encodes key fermentation genes. Virology 518, 423–433 (2018).
Yutin, N. & Koonin, E. V. Proteorhodopsin genes in giant viruses. Biol. Direct 7, 34 (2012).
Pushkarev, A. et al. A distinct abundant group of microbial rhodopsins discovered using functional metagenomics. Nature 558, 595–599 (2018).
Sharma, A. K., Spudich, J. L. & Doolittle, W. F. Microbial rhodopsins: functional versatility and genetic mobility. Trends Microbiol. 14, 463–469 (2006).
Fuhrman, J. A., Schwalbach, M. S. & Stingl, U. Proteorhodopsins: an array of physiological roles? Nat. Rev. Microbiol. 6, 488–494 (2008).
Shah, V., Chang, B. X. & Morris, R. M. Cultivation of a chemoautotroph from the SUP05 clade of marine bacteria that produces nitrite and consumes ammonium. ISME J. 11, 263–271 (2017).
Anantharaman, K. et al. Sulfur oxidation genes in diverse deep-sea viruses. Science 344, 757–760 (2014).
Roux, S. et al. Ecology and evolution of viruses infecting uncultivated SUP05 bacteria as revealed by single-cell- and meta-genomics. eLife 3, e03125 (2014).
Roux, S. et al. Ecogenomics and potential biogeochemical impacts of globally abundant ocean viruses. Nature 537, 689–693 (2016).
Ahlgren, N. A., Fuchsman, C. A., Rocap, G. & Fuhrman, J. A. Discovery of several novel, widespread, and ecologically distinct marine Thaumarchaeota viruses that encode amoC nitrification genes. ISME J. 13, 618–631 (2018).
Clokie, M. R. J. & Mann, N. H. Marine cyanophages and light. Environ. Microbiol. 8, 2074–2082 (2006).
Ni, T. & Zeng, Q. Diel infection of cyanobacteria by cyanophages. Front. Mar. Sci. 2, 123 (2016).
Cseke, C. S. & Farkas, G. L. Effect of light on the attachment of cyanophage AS-1 to Anacystis nidulans. J. Bacteriol. 137, 667–669 (1979).
Kao, C. C., Green, S., Stein, B. & Golden, S. S. Diel infection of a cyanobacterium by a contractile bacteriophage. Appl. Environ. Microbiol. 71, 4276–4279 (2005).
Jia, Y., Shan, J., Millard, A., Clokie, M. R. J. & Mann, N. H. Light-dependent adsorption of photosynthetic cyanophages to Synechococcus sp. WH7803. FEMS Microbiol. Lett. 310, 120–126 (2010).
Reimers, A.-M., Knoop, H., Bockmayr, A. & Steuer, R. Cellular trade-offs and optimal resource allocation during cyanobacterial diurnal growth. Proc. Natl Acad. Sci. USA 114, E6457–E6465 (2017).
Sheyn, U., Rosenwasser, S., Ben-Dor, S., Porat, Z. & Vardi, A. Modulation of host ROS metabolism is essential for viral infection of a bloom-forming coccolithophore in the ocean. ISME J. 10, 1742–1754 (2016).
Suttle, C. A. & Chen, F. Mechanisms and rates of decay of marine viruses in seawater. Appl. Environ. Microbiol. 58, 3721–3729 (1992).
Luo, E., Aylward, F. O., Mende, D. R. & Delong, E. F. Bacteriophage distributions and temporal variability in the ocean’s interior. mBio 8, e01903–e01917 (2017).
Kimura, S. et al. Diurnal infection patterns and impact of Microcystis cyanophages in a Japanese pond. Appl. Environ. Microbiol. 78, 5805–5811 (2012).
Yoshida, T. et al. Locality and diel cycling of viral production revealed by a 24 h time course cross-omics analysis in a coastal region of Japan. ISME J. 12, 1287–1295 (2018).
Liu, R., Liu, Y., Chen, Y., Zhan, Y. & Zeng, Q. Cyanobacterial viruses exhibit diurnal rhythms during infection. Proc. Natl Acad. Sci. USA 116, 14077–14082 (2019). This paper shows distinct diel-dependent life history traits in three Prochlorococcus phages, and that rhythmic phage transcription is linked to the photosynthetic activity of the host.
Bremer, H. et al. Escherichia Coli and Salmonella: Cellular and Molecular Biology 2nd edn Vol. 2 (eds Neidhardt, F. C. et al.) 1553–1569 (ASM Press, 1996).
Kirchman, D. L. Processes in Microbial Ecology 19–34 (Oxford Univ. Press, 2012).
Jover, L. F., Effler, T. C., Buchan, A., Wilhelm, S. W. & Weitz, J. S. The elemental composition of virus particles: implications for marine biogeochemical cycles. Nat. Rev. Microbiol. 12, 519–528 (2014).
Clasen, J. L. & Elser, J. J. The effect of host Chlorella NC64A carbon:phosphorus ratio on the production of Paramecium bursaria Chlorella Virus-1. Freshw. Biol. 52, 112–122 (2007).
Brown, C. M., Lawrence, J. E. & Campbell, D. A. Are phytoplankton population density maxima predictable through analysis of host and viral genomic DNA content? J. Mar. Biol. Assoc. UK 86, 491–498 (2006).
Edwards, K. F. & Steward, G. F. Host traits drive viral life histories across phytoplankton viruses. Am. Nat. 191, 566–581 (2018).
Wikner, J., Vallino, J. J., Steward, G. F., Smith, D. C. & Azam, F. Nucleic acids from the host bacterium as a major source of nucleotides for three marine bacteriophages. FEMS Microbiol. Ecol. 12, 237–248 (1993).
Kozloff, L. M. & Putnam, F. W. Biochemical studies of virus reproduction: III. The origin of virus phosphorus in the Escherichia coli T6 bacteriophage system. J. Biol. Chem. 182, 229–242 (1950).
Pasulka, A. L. et al. Interrogating marine virus–host interactions and elemental transfer with BONCAT and nanoSIMS-based methods. Environ. Microbiol. 20, 671–692 (2018).
Stent, G. S. & Maaløe, O. Radioactive phosphorus tracer studies on the reproduction of T4 bacteriophage. Biochim. Biophys. Acta 10, 55–69 (1953).
Sullivan, M. B. et al. Genomic analysis of oceanic cyanobacterial myoviruses compared with T4-like myoviruses from diverse hosts and environments. Environ. Microbiol. 12, 3035–3056 (2010).
Zeng, Q. & Chisholm, S. W. Marine viruses exploit their host’s two-component regulatory system in response to resource limitation. Curr. Biol. 22, 124–128 (2012).
Tetu, S. G. et al. Microarray analysis of phosphate regulation in the marine cyanobacterium Synechococcus sp. WH8102. ISME J. 3, 835–849 (2009).
Kelly, L., Ding, H., Huang, K. H., Osburne, M. S. & Chisholm, S. W. Genetic diversity in cultured and wild marine cyanomyoviruses reveals phosphorus stress as a strong selective agent. ISME J. 7, 1827–1841 (2013).
Bertilsson, S., Berglund, O., Karl, D. M. & Chisholm, S. W. Elemental composition of marine Prochlorococcus and Synechococcus: implications for the ecological stoichiometry of the sea. Limnol. Oceanogr. 48, 1721–1731 (2003).
Martiny, A. C., Coleman, M. L. & Chisholm, S. W. Phosphate acquisition genes in Prochlorococcus ecotypes: evidence for genome-wide adaptation. Proc. Natl Acad. Sci. USA 103, 12552–12557 (2006).
Wilson, W. H. et al. Complete genome sequence and lytic phase transcription profile of a Coccolithovirus. Science 309, 1090–1092 (2005).
Monier, A. et al. Host-derived viral transporter protein for nitrogen uptake in infected marine phytoplankton. Proc. Natl Acad. Sci. USA 114, E7489–E7498 (2017). This study reports the first nitrogen transport gene in an algal virus isolate and shows that it enables uptake of ammonium as well as organic nitrogen substrates.
Monier, A. et al. Phosphate transporters in marine phytoplankton and their viruses: cross-domain commonalities in viral–host gene exchanges. Environ. Microbiol. 14, 162–176 (2012).
Grossman, A. R., Schaefer, M. R., Chiang, G. G. & Collier, J. L. The phycobilisome, a light-harvesting complex responsive to environmental conditions. Microbiol. Rev. 57, 725–749 (1993).
Gao, E.-B., Gui, J.-F. & Zhang, Q.-Y. A novel cyanophage with a cyanobacterial nonbleaching protein A gene in the genome. J. Virol. 86, 236–245 (2012).
Ou, T., Gao, X. C., Li, S. H. & Zhang, Q. Y. Genome analysis and gene nblA identification of Microcystis aeruginosa myovirus (MaMV-DC) reveal the evidence for horizontal gene transfer events between cyanomyovirus and host. J. Gen. Virol. 96, 3681–3697 (2015).
Nadel, O. et al. Uncultured marine cyanophages encode for active NblA, phycobilisome proteolysis adaptor protein. Preprint at bioRxiv https://doi.org/10.1101/494369 (2018).
Ma, X., Coleman, M. L. & Waldbauer, J. R. Distinct molecular signatures in dissolved organic matter produced by viral lysis of marine cyanobacteria. Environ. Microbiol. 20, 3001–3011 (2018).
Sañudo-Wilhelmy, S. A. et al. Multiple B-vitamin depletion in large areas of the coastal ocean. Proc. Natl Acad. Sci. USA 109, 14041–14045 (2012).
Croft, M. T., Lawrence, A. D., Raux-Deery, E., Warren, M. J. & Smith, A. G. Algae acquire vitamin B12 through a symbiotic relationship with bacteria. Nature 438, 90–93 (2005).
Heal, K. R. et al. Two distinct pools of B12 analogs reveal community interdependencies in the ocean. Proc. Natl Acad. Sci. USA 114, 364–369 (2017).
López-Pérez, M., Haro-Moreno, J. M., de la Torre, J. R. & Rodriguez-Valera, F. Novel caudovirales associated with Marine Group I Thaumarchaeota assembled from metagenomes. Environ. Microbiol. 21, 1980–1988 (2019).
Ignacio-Espinoza, J. C. & Sullivan, M. B. Phylogenomics of T4 cyanophages: lateral gene transfer in the ‘core’ and origins of host genes. Environ. Microbiol. 14, 2113–2126 (2012).
Moore, J. K., Doney, S. C. & Lindsay, K. Upper ocean ecosystem dynamics and iron cycling in a global three-dimensional model. Glob. Biogeochem. Cycles 18, GB4028 (2004).
Bonnain, C., Breitbart, M. & Buck, K. N. The ferrojan horse hypothesis: iron–virus interactions in the ocean. Front. Mar. Sci. 3, 82 (2016).
Gledhill, M. et al. Effect of metals on the lytic cycle of the coccolithovirus, EhV86. Front. Microbiol. 3, 155 (2012).
Gasper, R. et al. Distinct features of cyanophage-encoded T-type phycobiliprotein lyase ΦCpeT: the role of auxiliary metabolic genes. J. Biol. Chem. 292, 3089–3098 (2017).
Nissimov, J. I. et al. Biochemical diversity of glycosphingolipid biosynthesis as a driver of Coccolithovirus competitive ecology. Environ. Microbiol. 21, 2182–2197 (2019).
Deng, L. et al. Grazing of heterotrophic flagellates on viruses is driven by feeding behaviour. Environ. Microbiol. Rep. 6, 325–330 (2014).
Baltar, F. Watch out for the ‘living dead’: cell-free enzymes and their fate. Front. Microbiol. 8, 2438 (2018).
Malits, A., Christaki, U., Obernosterer, I. & Weinbauer, M. G. Enhanced viral production and virus-mediated mortality of bacterioplankton in a natural iron-fertilized bloom event above the kerguelen plateau. Biogeosciences 11, 6841–6853 (2014).
Motegi, C. et al. Viral control of bacterial growth efficiency in marine pelagic environments. Limnol. Oceanogr. 54, 1901–1910 (2009).
Brum, J. R., Hurwitz, B. L., Schofield, O., Ducklow, H. W. & Sullivan, M. B. Seasonal time bombs: dominant temperate viruses affect southern ocean microbial dynamics. ISME J. 10, 437–449 (2016).
Weitz, J. S. et al. A multitrophic model to quantify the effects of marine viruses on microbial food webs and ecosystem processes. ISME J. 9, 1352–1364 (2015).
Talmy, D. et al. An empirical model of carbon flow through marine viruses and microzooplankton grazers. Environ. Microbiol. 21, 2171–2181 (2019). Using an empirically parameterized model constrained by estimates of prey, predator and viral life history traits, this study calculates carbon flows from primary producers to viruses, grazers and lysates in a marine ecosystem.
Lønborg, C., Middelboe, M. & Brussaard, C. P. D. Viral lysis of Micromonas pusilla: impacts on dissolved organic matter production and composition. Biogeochemistry 116, 231–240 (2013).
Lelchat, F. et al. Viral degradation of marine bacterial exopolysaccharides. FEMS Microbiol. Ecol. 95, fiz079 (2019).
Middelboe, M. & Jørgensen, N. O. G. Viral lysis of bacteria: an important source of dissolved amino acids and cell wall compounds. J. Mar. Biol. Assoc. UK 86, 605–612 (2006).
Yamada, Y., Tomaru, Y., Fukuda, H. & Nagata, T. Aggregate formation during the viral lysis of a marine diatom. Front. Mar. Sci. 5, 167 (2018).
Shelford, E. J., Middelboe, M., Møller, E. F. & Suttle, C. A. Virus-driven nitrogen cycling enhances phytoplankton growth. Aquat. Microb. Ecol. 66, 41–46 (2012).
Poorvin, L., Rinta-Kanto, J. M., Hutchins, D. A. & Wilhelm, S. W. Viral release of iron and its bioavailability to marine plankton. Limnol. Oceanogr. 49, 1734–1741 (2004).
Gobler, C. J., Hutchins, D. A., Fisher, N. S., Cosper, E. M. & Sanudo-Wilhelmy, S. A. Release and bioavailability of C, N, P, Se, and Fe following viral lysis of a marine chrysophyte. Limnol. Oceanogr. 42, 1492–1504 (1997).
Middelboe, M., Jørgensen, N. O. G. & Kroer, N. Effects of viruses on nutrient turnover and growth efficiency of noninfected marine bacterioplankton. Appl. Environ. Microbiol. 62, 1991–1997 (1996).
Jiao, N. et al. Microbial production of recalcitrant dissolved organic matter: long-term carbon storage in the global ocean. Nat. Rev. Microbiol. 8, 593–599 (2010).
Moreno, A. R. & Martiny, A. C. Ecological stoichiometry of ocean plankton. Ann. Rev. Mar. Sci. 10, 43–69 (2018).
Sullivan, M. B., Weitz, J. S. & Wilhelm, S. Viral ecology comes of age. Environ. Microbiol. Rep. 9, 33–35 (2017).
Weinbauer, M. G. Ecology of prokaryotic viruses. FEMS Microbiol. Rev. 28, 127–181 (2004).
Laber, C. P. et al. Coccolithovirus facilitation of carbon export in the North Atlantic. Nat. Microbiol. 3, 537–547 (2018). This field study marshals an array of evidence to provide some of the first direct measurements of the effects of viral infection on large-scale carbon export in a natural marine ecosystem.
Johns, C. T. et al. The mutual interplay between calcification and coccolithovirus infection. Environ. Microbiol. 21, 1896–1915 (2019).
Evans, C. & Wilson, W. H. Preferential grazing of Oxyrrhis marina on virus-infected Emiliania huxleyi. Limnol. Oceanogr. 53, 2035–2040 (2008).
Vermont, A. I. et al. Virus infection of Emiliania huxleyi deters grazing by the copepod Acartia tonsa. J. Plankton Res. 38, 1194–1205 (2016).
Frada, M. J. et al. Zooplankton may serve as transmission vectors for viruses infecting algal blooms in the ocean. Curr. Biol. 24, 2592–2597 (2014).
Lawrence, J. et al. Viruses on the menu: the appendicularian Oikopleura dioica efficiently removes viruses from seawater. Limnol. Oceanogr. 63, S244–S253 (2018).
Gonzalez, J. M. & Suttle, C. A. Grazing by marine nanoflagellates on viruses and virus-sized particles: ingestion and digestion. Mar. Ecol. Prog. Ser. 94, 1–10 (1993).
Record, N. R., Talmy, D. & Våge, S. Quantifying tradeoffs for marine viruses. Front. Mar. Sci. 3, 251 (2016).
Mateus, M. D. Bridging the gap between knowing and modeling viruses in marine systems — an upcoming frontier. Front. Mar. Sci. 3, 284 (2017).
Intergovernmental Panel on Climate Change. Climate Change 2014: synthesis report. Contribution of Working Groups I, II and III to the fifth assessment report of the Intergovernmental Panel on Climate Change (IPCC, 2014).
Stocker, T. F. et al. Climate Change 2013 — The Physical Science Basis (Cambridge Univ. Press, 2014).
Stock, C. A., Dunne, J. P. & John, J. G. Global-scale carbon and energy flows through the marine planktonic food web: an analysis with a coupled physical–biological model. Prog. Oceanogr. 120, 1–28 (2014).
Aumont, O., Ethé, C., Tagliabue, A., Bopp, L. & Gehlen, M. PISCES-v2: an ocean biogeochemical model for carbon and ecosystem studies. Geosci. Model. Dev. 8, 2465–2513 (2015).
Danovaro, R. et al. Marine viruses and global climate change. FEMS Microbiol. Rev. 35, 993–1034 (2011).
Nissimov, J. I., Napier, J. A., Allen, M. J. & Kimmance, S. A. Intragenus competition between coccolithoviruses: an insight on how a select few can come to dominate many. Environ. Microbiol. 18, 133–145 (2016).
Zimmerman, A. E. et al. Closely related viruses of the marine picoeukaryotic alga Ostreococcus lucimarinus exhibit different ecological strategies. Environ. Microbiol. 21, 2148–2170 (2019).
Martiny, A. C. et al. Strong latitudinal patterns in the elemental ratios of marine plankton and organic matter. Nat. Geosci. 6, 279–283 (2013).
Weber, T. S. & Deutsch, C. Ocean nutrient ratios governed by plankton biogeography. Nature 467, 550–554 (2010).
Weitz, J. S., Li, G., Gulbudak, H., Cortez, M. H. & Whitaker, R. J. Viral invasion fitness across a continuum from lysis to latency. Virus Evol. 5, vez006 (2019).
Mai-Prochnow, A. et al. ‘Big things in small packages: the genetics of filamentous phage and effects on fitness of their host’. FEMS Microbiol. Rev. 39, 465–487 (2015).
Brüssow, H., Canchaya, C. & Hardt, W.-D. Phages and the evolution of bacterial pathogens: from genomic rearrangements to lysogenic conversion. Microbiol. Mol. Biol. Rev. 68, 560–602 (2004).
Obeng, N., Pratama, A. A. & Elsas, J. D. van. The significance of mutualistic phages for bacterial ecology and evolution. Trends Microbiol. 24, 440–449 (2016).
Feiner, R. et al. A new perspective on lysogeny: prophages as active regulatory switches of bacteria. Nat. Rev. Microbiol. 13, 641–650 (2015).
Choua, M. & Bonachela, J. A. Ecological and evolutionary consequences of viral plasticity. Am. Nat. 193, 346–358 (2019).
Dang, V. T., Howard-Varona, C., Schwenck, S. & Sullivan, M. B. Variably lytic infection dynamics of large Bacteroidetes podovirus phi38:1 against two Cellulophaga baltica host strains. Environ. Microbiol. 17, 4659–4671 (2015).
Bryan, D., El-Shibiny, A., Hobbs, Z., Porter, J. & Kutter, E. M. Bacteriophage T4 infection of stationary phase E. coli: life after log from a phage perspective. Front. Microbiol. 7, 1391 (2016).
Mackinder, L. C. M. et al. A unicellular algal virus, Emiliania huxleyi virus 86, exploits an animal-like infection strategy. J. Gen. Virol. 90, 2306–2316 (2009).
Thomas, R. et al. Acquisition and maintenance of resistance to viruses in eukaryotic phytoplankton populations. Environ. Microbiol. 13, 1412–1420 (2011).
Adams, M. H. Bacteriophages (Interscience, 1959).
Hyman, P. & Abedon, S. T. in Bacteriophages. Methods and Protocols, Volume 1: Isolation, Characterization, and Interaction (eds Clokie, M. R. J. & Kropinski, A. M.) 175–202 (Humana Press, 2009).
Abedon, S. T. Phage therapy dosing: the problem(s) with multiplicity of infection (MOI). Bacteriophage 6, e1220348 (2016).
Mayer, J. A. & Taylor, F. J. R. A virus which lyses the marine nanoflagellate Micromonas pusilla. Nature 281, 299–301 (1979).
Brum, J. R. & Sullivan, M. B. Rising to the challenge: accelerated pace of discovery transforms marine virology. Nat. Rev. Microbiol. 13, 147–159 (2015).
Reistetter, E. N. et al. Effects of phosphorus starvation versus limitation on the marine cyanobacterium Prochlorococcus MED4 II: gene expression. Environ. Microbiol. 15, 2129–2143 (2013).
Guo, J. et al. Specialized proteomic responses and an ancient photoprotection mechanism sustain marine green algal growth during phosphate limitation. Nat. Microbiol. 3, 781–790 (2018).
Gerringa, L. J. A., de Baar, H. J. W. & Timmermans, K. R. A comparison of iron limitation of phytoplankton in natural oceanic waters and laboratory media conditioned with EDTA. Mar. Chem. 68, 335–346 (2000).
Mistry, B. A., D’Orsogna, M. R. & Chou, T. The effects of statistical multiplicity of infection on virus quantification and infectivity assays. Biophys. J. 114, 2974–2985 (2018).
Kirzner, S., Barak, E. & Lindell, D. Variability in progeny production and virulence of cyanophages determined at the single-cell level. Environ. Microbiol. Rep. 8, 605–613 (2016).
Cheng, Y. S., Labavitch, J. & VanderGheynst, J. S. Organic and inorganic nitrogen impact Chlorella variabilis productivity and host quality for viral production and cell lysis. Appl. Biochem. Biotechnol. 176, 467–479 (2015).
Maat, D. S., Crawfurd, K. J., Timmermans, K. R. & Brussaard, C. P. D. Elevated CO2 and phosphate limitation favor Micromonas pusilla through stimulated growth and reduced viral impact. Appl. Environ. Microbiol. 80, 3119–3127 (2014).
Acknowledgements
This work was supported by the Gordon & Betty Moore Foundation Marine Microbiology Initiative (Award 3305). Additional support was provided by the National Science Foundation Division of Ocean Sciences (NSF-OCE) (Awards 1536989 and 1829831 to M.B.S.), the Simons Foundation (Awards 32910 to S.J. and 402971 to J.R.W.) and the Gordon & Betty Moore Foundation (Awards 3788 to A.Z.W. and 3790 to M.B.S.).
Author information
Authors and Affiliations
Contributions
All authors contributed to the conception, research, writing and editing of the article.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information
Nature Reviews Microbiology thanks K. Bidle and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Glossary
- Transparent exopolymer particle
-
(TEP). A sticky, gel-like particle consisting predominantly of acidic polysaccharides that originate from microorganisms and can enhance the aggregation of non-sticky particles in marine and aquatic ecosystems.
- Core reaction centre
-
A membrane complex of several proteins, pigments and other cofactors that performs the principal energy conversion reactions of photosynthesis, capturing light energy and converting it into redox potential energy for ATP synthesis and reducing power for reduction of CO2; also known as the photosynthetic reaction centre.
- Phycobilin
-
Photosynthetic pigments found in cyanobacteria and the chloroplasts of red algae and glaucophytes that aid in absorption of light energy, particularly at wavelengths that are not well absorbed by chlorophylls or carotenoids.
- Euphotic zone
-
The uppermost layer of water in a lake or ocean characterized by enough sunlight to support photosynthetic carbon fixation.
Rights and permissions
About this article
Cite this article
Zimmerman, A.E., Howard-Varona, C., Needham, D.M. et al. Metabolic and biogeochemical consequences of viral infection in aquatic ecosystems. Nat Rev Microbiol 18, 21–34 (2020). https://doi.org/10.1038/s41579-019-0270-x
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41579-019-0270-x
This article is cited by
-
Exploring virus-host-environment interactions in a chemotrophic-based underground estuary
Environmental Microbiome (2024)
-
Hidden diversity and potential ecological function of phosphorus acquisition genes in widespread terrestrial bacteriophages
Nature Communications (2024)
-
Algal blooms in the ocean: hot spots for chemically mediated microbial interactions
Nature Reviews Microbiology (2024)
-
COBRA improves the completeness and contiguity of viral genomes assembled from metagenomes
Nature Microbiology (2024)
-
Viral lysing can alleviate microbial nutrient limitations and accumulate recalcitrant dissolved organic matter components in soil
The ISME Journal (2023)