Abstract
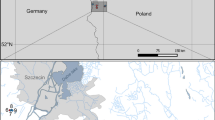
We have studied the distribution and community composition of denitrifying bacteria in the stratified water column and at the sediment–water interface in lakes Plußsee and Schöhsee, and a near-shore site in the Baltic Sea in Germany. Although environmental changes induced by the stratification of the water column in marine environments are known to affect specific populations of denitrifying bacteria, little information is available for stratified freshwater lakes and brackish water. The aim of the present study was to fill this gap and to demonstrate specific distribution patterns of denitrifying bacteria in specific aquatic habitats using two functional markers for the nitrite reductase (nirK and nirS genes) as a proxy for the communities. The leading question to be answered was whether communities containing the genes nirK and nirS have similar, identical, or different distribution patterns, and occupy the same or different ecological niches. The genes nirK and nirS were analyzed by PCR amplification with specific primers followed by terminal restriction fragment length polymorphism (T-RFLP) and by cloning and sequence analysis. Overall, nirS-denitrifiers were more diverse than nirK-denitrifiers. Denitrifying communities in sediments were clearly different from those in the water column in all aquatic systems, regardless of the gene analyzed. A differential distribution of denitrifying assemblages was observed for each particular site. In the Baltic Sea and Lake Plußsee, nirK-denitrifiers were more diverse throughout the water column, while nirS-denitrifiers were more diverse in the sediment. In Lake Schöhsee, nirS-denitrifiers showed high diversity across the whole water body. Habitat-specific clusters of nirS sequences were observed for the freshwater lakes, while nirK sequences from both freshwater lakes and the Baltic Sea were found in common phylogenetic clusters. These results demonstrated differences in the distribution of bacteria containing nirS and those containing nirK indicating that both types of denitrifiers apparently occupy different ecological niches.





Similar content being viewed by others
References
Aberle N, Wiltshire KH (2006) Seasonality and diversity patterns of microphytobenthos in a mesotrophic lake. Arch Hydrobiol 167:447–465
Altschul SF, Madden TL, Schäffer AA, Zhang J, Zhang ZWM, Lipman DJ (1997) Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res 25:3389–3402
Avrahami S, Conrad R, Braker G (2002) Effect of soil ammonium concentration on N2O release and on the community structure of ammonia oxidizers and denitrifiers. Appl Environ Microbiol 68:5685–5692
Braker G, Zhou J, Wu L, Devol AH, Tiedje JM (2000) Nitrite reductase genes (nirK and nirS) as functional markers to investigate diversity of denitrifying bacteria in Pacific Northwest marine sediment community. Appl Environ Microbiol 66:2096–2104
Brettar I, Moore ERB, Höfle MG (2001) Phylogeny and abundance of novel denitrifying bacteria isolated from the water column of the central Baltic Sea. Microb Ecol 42:295–305
Castro-González M, Braker G, Farías L, Ulloa O (2005) Communities of nirS-type denitrifiers in the water column of the oxygen minimum zone in the eastern South Pacific. Environ Microbiol 7:1298–1306
Coyne MS, Arunakumari A, Averill BA, Tiedje JM (1989) Immunological identification and distribution of dissimilatory heme cd1 and non-heme copper nitrite reductases in denitrifying bacteria. Appl Environ Microbiol 55:2924–2931
Eckert W, Imberger J, Saggio A (2002) Biogeochemical response to physical forcing in the water column of a warm monomictic lake. Biogeochemistry 61:291–307
Falk S, Hannig M, Gliesche C, Wardenga R, Koster M, Jürgens K, Braker G (2007) nirS-containing denitrifiers communities in the water column and sediment of the Baltic Sea. Biogeosciences 4:255–268
Hallin S, Lindgren PE (1999) PCR detection of genes encoding nitrite reductase in denitrifying bacteria. Appl Environ Microbiol 65
Hannig M, Braker G, Dppner J, Jürgens K (2006) Linking denitrifier community structure and prevalent biogeochemical parameters in the pelagial of the central Baltic Sea Proper (Baltic Sea). FEMS Microbiol Ecol 57:260–271
Jayakumar DA, Francis CA, Naqvi SWA, Ward BB (2004) Diversity of nitrite reductase genes (nirS) in the denitrifying water column of the coastal Arabian Sea. Aquat Microb Ecol 34:69–78
Junier P, Junier T, Witzel KP (2008a) TRiFLe, a program for in silico terminal restriction fragment length polymorphism analysis with user-defined sequence sets. Appl Environ Microbiol 74:6452–6456
Junier P, Kim O-S, Witzel K-P, Imhoff JF, Hadas O (2008b) Habitat-partitioning of denitrifying bacterial communities carrying nirS/nirK genes in the stratified water column of Lake Kinneret, Israel. Aquatic Microb Ecol 51:129–140
Kim OS, Junier P, Imhoff JF, Witzel KP (2008) Comparative analysis of ammonia monooxygenase (amoA) genes in the water column and sediment-water interface of two lakes and the Baltic Sea. FEMS Microbiol Ecol 66:367–378
Knowles R (1982) Denitrification. Microbio Rev 46:43–70
Liu X, Tiquia SM, Holguin G, Wu L, Nold SC, Devol AH, Luo K, Palumbo AV, Tiedje JM, Zhou J (2003) Molecular diversity of denitrifying genes in continental margin sediments within the oxygen-deficient zone off the Pacific coast of Mexico. Appl Environ Microbiol 69:3549–3560
Michotey V, Mejean V, Bonin P (2000) Comparison of methods for quantification of cytochrome cd1-denitrifying bacteria in environmental marine samples. Appl Environ Microbiol 66:1564–1571
Nogales B, Timmis KN, Nedwell DB, Osborn AM (2002) Detection and diversity of expressed denitrification genes in estuarine sediments after reverse transcription-PCR amplification from mRNA. Appl Environ Microbiol 68:5017–5025
Oakley BB, Francis CA, Roberts KJ, Fuchsman CA, Srinivasan S, Staley JT (2007) Analysis of nitrite reductase (nirK and nirS) genes and cultivation reveal depauperate community of denitrifying bacteria in the Black Sea suboxic zone. Environ Microbiol 9:118–130
Overbeck J, Chróst RJ (1994) Microbial ecology of Lake Plußsee. Ecol Studies 105:1–44
Philippot L (2002) Denitrifying genes in bacterial and archaeal genomes. Biochim Biophys Acta 1577:355–376
Priemé A, Braker G, Tiedje JM (2002) Diversity of nitrite reductase (nirK and nirS) gene fragments in forested upland and wetland soils. Appl Environ Microbiol 68:1893–1900
Rai H, Arts MT, Wainman BC, Dockal N, Krambeck HJ (1997) Lipid production in natural phytoplankton communities in a small freshwater Baltic lake, Lake Schöhsee, Germany. Freshw Biol 38:581–590
Rösch C, Mergel A, Bothe H (2002) Biodiversity of denitrifying and dinitrogen-fixing bacteria in an acid forest soil. Appl Environ Microbiol 68:3818–3829
Santoro AE, Boehm AB, Francis CA (2006) Denitrifier community composition along a nitrate and salinity gradient in a coastal aquifer. Appl Environ Microbiol 72:2102–2109
Satoh T, Hoshino Y, Kitamura H (1976) Rhodopseudomonas sphaeroides forma sp. denitrificans, a denitrifying strain as a subspecies of Rhodopseudomonas sphaeroides. Arch Microbiol 108:265–269
Schäfer H, McDonald IR, Nightingale PD, Murrell JC (2005) Evidence for the presence of a CmuA methyltransferase pathway in novel marine methyl halide-oxidizing bacteria. Environ Microbiol 7:839–852
Schloss PD, Handelsman J (2005) Introducing DOTUR, a computer program for defining operational taxonomic units and estimating species richness. Appl Environ Microbiol 71:1501–1506
Sharma S, Aneja MK, Mayer J, Munch JC, Schloter M (2005) Diversity of transcripts of nitrite reductase genes (nirK and nirS) in rhizospheres of grain legumes. Appl Environ Microbiol 71:2001–2007
Taroncher-Oldenburg G, Griner EM, Francis CA, Ward BB (2003) Oligonucleotide microarray for the study of functional gene diversity in the nitrogen cycle in the environment. Appl Environ Microbiol 69:1159–1171
Throbäck IN, Enwall K, Jarvis Å, Hallin S (2004) Reassessing PCR primers targeting nirS, nirK and nosZ genes for community surveys of denitrifying bacteria with DGGE. FEMS Microbiol Ecol 49:401–417
Throbäck IN, Johansson M, Rosenquist M, Pell M, Hansson M, Hallin S (2007) Silver (Ag+) reduces denitrification and induces enrichment of novel nirK genotypes in soil. FEMS Microbiol Lett 270:189–194
Tuomainen JM, Hietanen S, Kuparinen J, Martikainen PJ, Servomaa K (2003) Baltic Sea cyanobacterial bloom contains denitrification and nitrification genes, but has negligible denitrification activity. FEMS Microbiol Ecol 45:83–96
Wolsing M, Priemé A (2004) Observation of high seasonal variation in community sturucture of denitrifying bacteria in arable soil receiving artificial fertilizer and cattle manure by determining T-RFLP of nir gene fragments. FEMS Microbiol Ecol 48:261–271
Yan T, Fields MW, Wu L, Zu Y, Tiedje JM, Zhou J (2003) Molecular diversity and characterization of nitrite reductase gene fragments (nirK and nirS) from nitrate- and uranium-contaminated groundwater. Environ Microbiol 5:13–24
Yoshie S, Noda N, Tsuneda S, Hirata A, Inamori Y (2004) Salinity decreases nitrite reductase gene diversity in denitrifying bacteria of wastewater treatment systems. Appl Environ Microbiol 70:3152–3157
Zumft WG (1997) Cell biology and molecular basis of denitrification. Microbiol Mol Biol Rev 61:533–616
Author information
Authors and Affiliations
Corresponding author
Additional information
Handling Editor: Piet Spaak.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Kim, OS., Imhoff, J.F., Witzel, KP. et al. Distribution of denitrifying bacterial communities in the stratified water column and sediment–water interface in two freshwater lakes and the Baltic Sea. Aquat Ecol 45, 99–112 (2011). https://doi.org/10.1007/s10452-010-9335-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10452-010-9335-7