Abstract
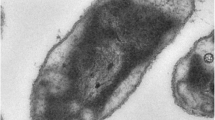
In a conserved culture of the purple sulfur bacterium Thiospirillum jenense DSM216T, cells of this species were easily recognized by cell morphology, large-size spirilla and visible flagellar tuft. The Tsp. jenense genome is 3.22 Mb in size and has a GC content of 48.7 mol%. It was readily identified as a member of the Chromatiaceae by the complement of proteins in its genome. A whole genome comparison clearly placed Tsp. jenense near Thiorhodovibrio and Rhabdochromatium species and somewhat more distant from Thiohalocapsa and Halochromatium species. This relationship was also found with the sequences of the photosynthetic reaction center protein PufM. The genome sequence supported important properties of this bacterium: the presence of ribulose-bisphosphate carboxylase and enzymes of the Calvin cycle of autotrophic carbon dioxide fixation but the absence of carboxysomes, an incomplete tricarboxylic acid cycle and the lack of malate dehydrogenase, the presence of a sulfur oxidation pathway including adenylylsulfate reductase (aprAB) but absence of assimilatory sulfate reduction, the presence of hydrogenase (hoxHMFYUFE), nitrogenase and a photosynthetic gene cluster (pufBALMC). The FixNOP type of cytochrome oxidase was notably lacking, which may be the reason that renders the cells highly sensitive to oxygen. Two minor phototrophic contaminants were found using metagenomic binning: one was identified as a strain of Rhodopseudomonas palustris and the second one has an average nucleotide identity of 82% to the nearest neighbor Rhodoferax antarcticus. It should be considered as a new species of this genus and Rhodoferax jenense is proposed as the name.






Similar content being viewed by others
References
Altschul SF, Madden TL, Schäffer AA, Zhang J, Zhang Z, Miller W, Lipman DJ (1997) Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res 25:3389–3402
Aziz RK, Bartels D, Best AA, DeJongh M, Disz T, Edwards RA, Fromsma K, Gerdes S, Glass EM, Kabul M, Meyer F, Olsen GJ, Olson R, Osterman AL, Overbeek RA, McNeil LK, Paarmann D, Paczian T, Parello B, Push GD, Reich C, Stevens R, Vassieva O, Vonstein V, Wilke A, Zagnitko O (2008) The RAST server: rapid annotations using subsystems technology. BMC Genomics 9(75):1
Badger MR, Bek EJ (2008) Multiple Rubisco forms in Proteobacteria: their functional significance in relation to CO2 acquisition by the CBB cycle. J Exp Bot 59:1525–1541
Briée C, Moreira D, López-García P (2007) Archaeal and bacterial community composition of sediment and plankton from a suboxic freshwater pond. Res Microbiol 158:213–227
Buder J (1915) Zur Kenntis des Thiospirillum jenense und seiner Reaktion auf Lichtreize. JB Bot 56:29–584
Buder J (1919) Zur Biologie des Bacteriopurpurins und der Purpurbakterien. JB Bot 58:525–628
Caldwell DE, Tiedje JM (1975) The structure of anaerobic bacterial communities in the hypolimnia of several Michigan lakes. Can J Microbiol 21:377–385
Ehrenberg CG (1838) Die Infusionstierchen als vollkommene Organismen. L. Voss-Verlag, Leipzig
Hungate RE (1969) A roll tube method for cultivation of strict anaerobes. In: Norris JR, Ribbons DW (eds) Methods in microbiology, vol 3B. Academic Press., NewYork, pp 117–132
Imhoff JF (2016) New dimensions in microbial ecology – Functional genes in studies to unravel the biodiversity and role of functional microbial groups in the environment. Microorganisms 4(2):19. https://doi.org/10.3390/microorganisms4020019
Imhoff JF, Meyer TE, Kyndt JA (2020) Genomic and genetic sequence information of strains assigned to the genus Rhodopseudomonas reveal the great heterogeneity of the group and identify strain Rhodopseudomonas palustris DSM 123T as the true type strain of this species. Int J Syst Evol Microbiol. https://doi.org/10.1099/ijsem.0.004077
Larkin MA, Blackshields G, Brown NP, Chenna R, McGettigan PA, McWilliam H, Valentin F, Wallace IM, Wilm A, Lopez R, Thompson JD, Gibson TJ, Higgins DG (2007) Clustal W and Clustal X version 2.0. Bioinformatics 23:2947–2948
Letunic I, Bork P (2019) Interactive tree of life (iTOL) v4: recent updates and new developments. Nucleic Acids Res 47:256–259. https://doi.org/10.1093/nar/gkz239
Mandel M, Leadbetter ER, Pfennig N, Trüper HG (1971) Deoxyribonucleic acid base compositions of phototrophic bacteria. Int J Syst Bacteriol 21:222–230
Neumann S, Wynen A, Trüper HG, Dahl C (2000) Characterization of the cys gene locus from Allochromatium vinosum indicates an unusual sulfate assimilation pathway. Mol Biol Rep 27:27–33
Parks DH, Imelfort M, Skennerton CT, Hugenholtz P, Tyson GW (2014) Assessing the quality of microbial genomes recovered from isolates, single cells, and metagenomes. Genome Res 25:1043–1055
Paterek JR, Paynter MJ (1988) Populations of anaerobic phototrophic bacteria in a Spartina alterniflora salt marsh. Appl Environ Microbiol 54:1360–1364
Pfennig N (1963) Thiospirillum jenense (Thiorhodaceae)— Lokomotion und phototaktisches Verhalten. IWF. https://doi.org/10.3203/IWF/E-678
Pfennig N (1993) Reflections of a microbiologist, or how to learn from the microbes. Annu Rev Microbiol 47:1–29
Perrière G, Gouy M (1996) WWW-Query: an on-line retrieval system for biological sequence banks. Biochimie 78:364–369
Preisig O, Zufferey R, Thöny-Meyer L, Appleby CA, Hennecke H (1996) A high-affinity cbb3-type cytochrome oxidase terminates the symbiosis-specific respiratory chain of Bradyrhizobium japonicum. J Bacteriol 178:1532–1538
Richter M, Rosselló-Móra R, Glöckner FO, Peplies J (2015) JSpeciesWS: a web server for prokaryotic species circumscription based on pairwise genome comparison. Bioinformatics 32:929-931. https://doi.org/10.1093/bioinformatics/btv681
Saitou N, Nei M (1987) The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4:406–425
Schlegel HG (1956) Vergleichende untersuchungen über die lichtempfindlichkeit einiger purpurbakterien. Arch Protistenkde 101:69–97
Schlegel HG, Pfennig N (1961) Die anreicherungskultur einiger schwefelpurpurbakterien. Arch Mikrobiol 38:1–39
Schmidt K (1963) Carotenoids in Thiorhodaceae. II. Carotenoid composition of Thiospirillum jenense Winogradsky and Chromatium vinosum Winogradsky. Arch Mikrobiol 46:127–137
Schmidt K, Pfennig N, Liaaen Jensen S (1965) Carotenoids of Thiorhodaceae. IV. The carotenoid composition of 25 pure isolates. Arch Mikrobiol 52:132–146
Stamatakis A, Hoover P, Rougemont JJSB (2008) A rapid bootstrap algorithm for the RAxML web servers. Syst Biol 57:758–771
Stamatakis AJB (2014) RAxML version 8: a tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics 30:1312–1313
Tabita FR, McFadden BA (1974) D-ribulose 1, 5-diphosphate carboxylase from Rhodospirillum rubrum. II. Quaternary structure, composition, catalytic and immunological properties. J Biol Chem 249:3459–3464
Tabita FR, Satagopan S, Hanson TE, Kreel NE, Scott SS (2008) Distinct form I, II, III, and IV rubisco proteins from the three kingdoms of life provide clues about rubisco evolution and structure/function relationships. J Experim Bot 59:1515–1524
Tank M, Thiel V, Imhoff JF (2009) Phylogenetic relationship of phototrophic purple sulfur bacteria according to pufL and pufM genes. Intern Microbiol 12:175–185
Tank M, Blümel M, Imhoff JF (2011) Communities of purple sulfur bacteria in a Baltic Sea coastal lagoon analyzed by pufLM gene libraries and the impact of temperature and NaCl concentration in experimental enrichment cultures. FEMS Microbiol Ecol 78:428–438
Thiel V, Tank M, Neulinger SC, Gehrmann L, Dorador C, Imhoff JF (2010) Unique communities of anoxygenic phototrophic bacteria in saline lakes of Salar de Atacama (Chile). Evidence for a new phylogenetic lineage of phototrophic Gammaproteobacteria from pufLM gene analyses. FEMS Microbiol Ecol 74:510–522
Van Niel CB (1932) On the morphology and physiology of the purple and green sulfur bacteria. Arch Mikrobiol 3:1–112
Van Niel CB (1955) The microbe as a whole. In: Waksman SA (ed) Perspectives and horizons in microbiology. Rutgers University Press, New Brunswick, NJ, pp 3–12
Wattam AR, Davis JJ, Assaf R, Boisvert S, Brettin T, Bun C, Conrad N, Dietrich EM, Disz T, Gabbard JL, Gerdes S, Henry CS, Kenyon RW, Machi D, Mao C, Nordberg EK, Olsen GJ, Murphy-Olson DE, Olson R, Overbeek R, Parrello B, Pusch GD, Shukla M, Vonstein V, Warren A, Xia F, Yoo H, Stevens RL (2017) Improvements to patric, the all-bacterial bioinformatics database and analysis resource center. Nucleic Acids Res 45:D535–D542
Weissgerber T, Zigann R, Bruce D, Chang Y, Detter JC, Han C, Hauser L, Jeffries CD, Land M, Munk AC, Tapia R, Dahl C (2011) Complete genome sequence of Allochromatium vinosum DSM 180T. Stand Genomic Sci 5:311–330
Weisburg WG, Barns SM, Pelletier DA, Lane DJ (1991) 16S ribosomal DNA amplification for phylogenetic study. J Bacteriol 173:697–703. https://doi.org/10.1128/jb.173.2.697-703.1991
Winogradsky SN (1888) Beiträge zur Morphologie und Physiologie der Schwefelbakterien, Leipzig, Arthur Felix, pp 1–120
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Communicated by Erko Stackebrandt.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Imhoff, J.F., Meyer, T.E. & Kyndt, J.A. The genome sequence of the giant phototrophic gammaproteobacterium Thiospirillum jenense gives insight into its physiological properties and phylogenetic relationships. Arch Microbiol 203, 97–105 (2021). https://doi.org/10.1007/s00203-020-02006-7
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00203-020-02006-7